New technique tracks individual protein movement on live cells

The piece of gold that Richard Taylor was thrilled to track down weighed less than a single bacterium. Taylor, a postdoctoral fellow at the Max Planck Institute, was working to follow individual nanogold-labeled molecules that move just nanometers, billionths of a meter. The resulting microscopy technique, developed under Professor Vahid Sandoghdar and colleagues, can follow proteins at microsecond speeds for long periods and will be presented today at the 64th Annual Meeting at the Biophysical Society in San Diego, California.
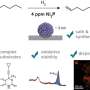
The microscope you may have used in high school biology is known as a brightfield microscope—it's the simplest microscopy technique. Light is transmitted through the sample and the magnifying lens, and you see the variations in density in the sample. But, if you're working to increase the sensitivity and see something smaller, brightfield reflects and scatters light, so some microscopy techniques add filters to eliminate the light scattering. Instead, Taylor and colleagues decided to take advantage of that scattered light. The light waves, reflected from the brightfield and scattered by gold particles used to label proteins, interfere with each other and the research team developed computational techniques to separate the desired signal from the rest. The method has been named interferometric scattering (iSCAT) microscopy.
"It's very sensitive, you can localize proteins very cleanly and precisely in three dimensions," Taylor explained. Compared to new microscopy techniques that create stunning images of cells, Taylor says, "ours is not quite as exotic, it is really a simple concept, the beauty is its simplicity." And unlike fluorescence microscopy, whose signal degrades over time, gold particles can be followed indefinitely.
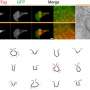
For the first test of the technique, Taylor and colleagues looked at gold-labeled proteins in solution. Then to try it in live cells, they chose a well-studied protein called epidermal growth factor receptor (EGFR), so they could confirm that their measurements were in line with all that was already known about the protein. Taylor says when he and his colleagues, who were all physicists, started looking at living cells, "we weren't ready for the amazing things we were about to see."
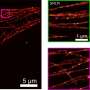
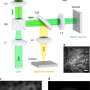
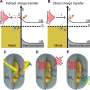
The dynamics of EGFR in cells astounded them and their biology collaborators—they watched as the protein diffused across the membrane, found its way into tapering membrane projections, and sank into pits to be internalized by the cell. Taylor said it reminded him of a "nano-rover" mapping the cell surface like a NASA vehicle on Mars. The movements that the computer tracked over long periods looked a bit like furious scribbles in two dimensions, but in three dimensions they resembled land topography.
EGFR is the only protein they've tracked so far, but in theory, they could track any cell surface protein, and may be able to track proteins within cells too. "The cell will scatter the signal, but that depends on the kind of cell, and where in the cell you're looking," Taylor said. They can also combine iSCAT with live cell fluorescence microscopy, which allows them to follow single proteins while also visualizing cell parts that might be influencing the way the proteins move, like the cell's scaffolding.
Taylor is excited to have the technique applied to other proteins, "we encourage scientists to use this microscopy—pick the protein you want to follow, and we'll show you how." He'll tell you exactly where to find your own tiny gold.
Provided by Biophysical Society