Researchers study protein ancestors to understand their role in growth
"Resurrecting" the ancestors of key proteins yields evolutionary insights into their role in human cells and in most cancers, a new study finds.
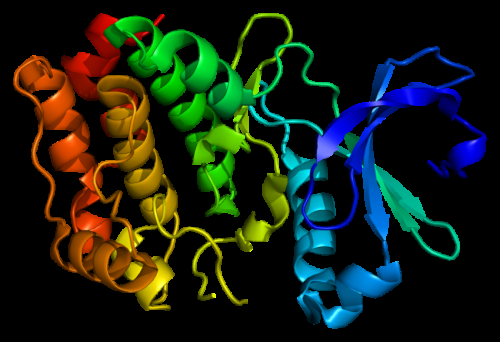
The study, published online August 13 in the journal eLife, revolves around the function of extracellular signal-regulated kinase (ERK), a protein that determines whether human cells divide and multiply as part of growth.
Led by researchers from NYU School of Medicine, the new study suggests that ERK was easily switched on in ancient one-celled creatures, but was carefully restrained as the first animals evolved 800 million years ago.
Specifically, the research team used computer analyses to determine the DNA sequences, and related protein structures, for ancient ERK relatives. By studying them, the researchers identified two key changes that likely brought ERK under more careful control across evolution.
The team also found that it was "surprisingly easy" to reverse those evolutionary changes and return ERK to its primitive, very active, ancestral state. This becomes important, the researchers say, because a similar process occurs as genetic mistakes convert normal cells into cancer. Modern-day ERK is part of "the most important signaling pathway in human cancers," along with the other protein switches that get stuck in the "on" position to cause abnormal growth.
"That ERK could continue to function while evolving reveals a flexibility that may explain how human cells evolved 500 different kinases that control all aspects of their biology—and why faulty kinases are so central to cancer growth," says senior study author Liam Holt, Ph.D., assistant professor in the Institute for Systems Genetics at NYU Langone Health. "Genetic changes are required for evolution to proceed, but many enzymes stop working when changes occur. ERK continues to signal."
Like all proteins, versions of ERK are assembled from combinations of molecular building blocks called amino acids. The growing variety of amino acid structures across the animal kingdom now catalogued in databases has made possible a new kind of analysis, say the authors. Researchers can make educated guesses about the amino acids occupying each position in the structures of their common ancestors.
Moving forward, the research team plans to resurrect a complete set of inferred ancestors for all proteins in ERK's class (kinases), and to screen drug candidates against them, says Holt. Their plan is based on the hypothesis that some mutations cause cancer because they return kinases to more ancient states that are not properly regulated.
To avert side effects and drug resistance, he says, future drugs might be designed to inhibit ancestor-like, cancer-causing kinase versions without turning off related, normal enzymes.
More information: Suk ho Hong et al, Learning from ancestors, eLife (2019). DOI: 10.7554/eLife.49976
Journal information: eLife
Provided by NYU Langone Health