'Semi-synthetic' bacteria churn out unnatural proteins
Synthetic biologists seek to create new life with forms and functions not seen in nature. Although scientists are a long way from making a completely artificial life form, they have made semi-synthetic organisms that have an expanded genetic code, allowing them to produce never-before-seen proteins. Now, researchers reporting in Journal of the American Chemical Society have optimized a semi-synthetic bacteria to efficiently produce proteins containing unnatural amino acids.
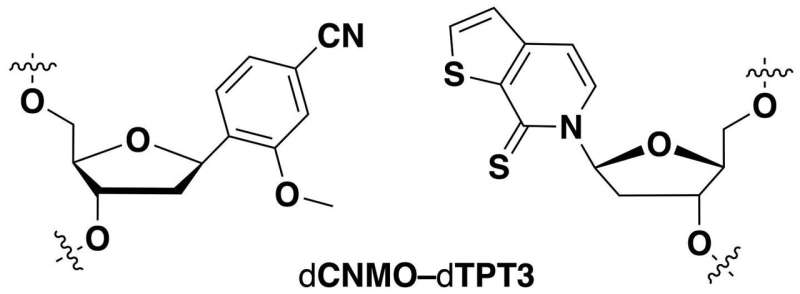
All of Earth's natural life forms store information using a four-letter genetic code consisting of the nucleotides deoxyadenosine (dA), deoxyguanosine (dG), deoxycytidine (dC), and deoxythymidine (dT). Within the DNA double helix, dA pairs with dT, and dG with dC, to form the "rungs" of the DNA ladder. Recently, researchers have made synthetic nucleotides that can pair up with each other. When they placed these unnatural nucleotides into genes, bacteria could replicate the DNA and convert the sequences into RNA and then proteins that contained unconventional amino acids. However, bacteria often cannot use these synthetic sequences as efficiently as the natural ones. Therefore, Lingjun Li, Floyd Romesberg and colleagues wanted to optimize the unnatural base pairs to improve protein production.
The researchers tested different combinations of unnatural base pairs in E. coli and observed which ones were replicated most efficiently and produced the highest levels of a protein. Some of the synthetic base pairs had been tested before, whereas others were new variations. The team then used these optimized base pairs to demonstrate, for the first time, a semi-synthetic organism that could make a protein containing multiple unnatural amino acids.
More information: Aaron W. Feldman et al, Optimization of Replication, Transcription, and Translation in a Semi-Synthetic Organism, Journal of the American Chemical Society (2019). DOI: 10.1021/jacs.9b02075
Journal information: Journal of the American Chemical Society
Provided by American Chemical Society