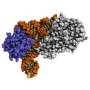
Expanding DNA's alphabet lets cells produce novel proteins
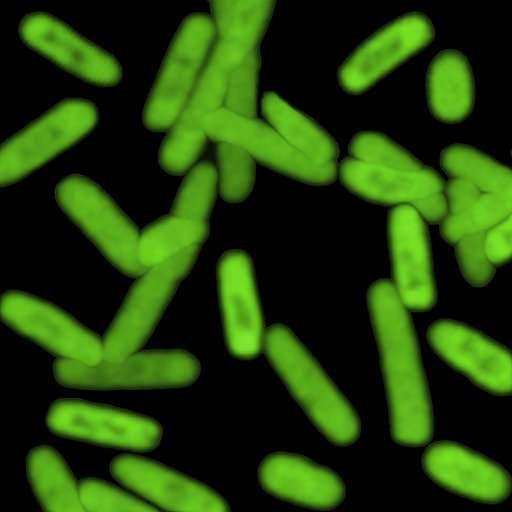
Scientists are expanding the genetic code of life, using man-made DNA to create a semi-synthetic strain of bacteria—and new research shows those altered microbes actually worked to produce proteins unlike those found in nature.
It's a step toward designer drug development.
One of the first lessons in high school biology: All life is made up of four DNA building blocks known by the letters A, T, C and G. Paired together, they form DNA's ladder-like rungs. Now there's a new rung on that ladder.
A team at The Scripps Research Institute in La Jolla, California, expanded the genetic alphabet, creating two artificial DNA "letters" called X and Y. A few years ago, the researchers brewed up a type of E. coli bacteria commonly used for lab research that contained both natural DNA and this new artificial base pair—storing extra genetic information inside cells.
The next challenge: Normal DNA contains the coding for cells to form proteins that do the work of life. Could cells carrying this weird genomic hybrid work the same way?
Sure enough, the altered cells glowed green as they produced a fluorescent protein containing unnatural amino acids, researchers reported Wednesday in the journal Nature.
"We can make proteins that are built of more things than they normally are," explained Scripps chemist Floyd Romesberg, who leads the project.
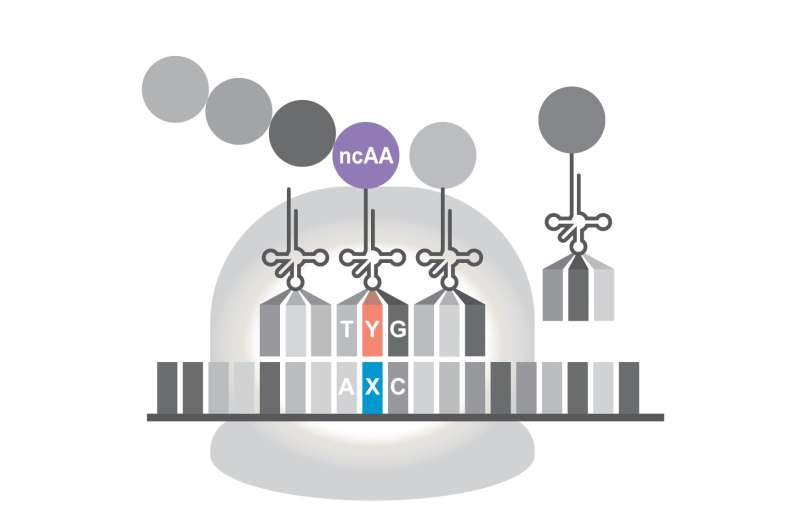
While programming the green germs offered evidence that the approach can work, eventually "we would like to get proteins that do new things," he said.
That's an ultimate goal in the field of synthetic biology—designing organisms that work differently from the way nature intended so scientists can harness them to create designer drugs, biofuels or a range of other products. Scripps' technology has been licensed by a biotech company Romesberg co-founded, Synthorx Inc., that aims to make novel protein-based drugs.
The new work traced the biological steps as the altered E. coli read the artificial genetic code and assembled the pieces for a new protein, with the same efficiency as if using normal DNA.
The result is a platform that offers a way to increase the diversity of proteins made inside living cells, said Jef Boeke, a synthetic biology researcher at New York University who wasn't involved in Scripps' work.
This bacterial strain was "modified in a really dramatic and unusual way at these positions in its genome," Boeke said. "And that's what makes it different from every other organism on the planet today."
More information: Yorke Zhang et al. A semi-synthetic organism that stores and retrieves increased genetic information, Nature (2017). DOI: 10.1038/nature24659
Journal information: Nature
© 2017 The Associated Press. All rights reserved.