Genome-wide rules of nucleosome phasing in drosophila

LMU researchers have, for the first time, systematically determined the positioning of the packing units of the fruit fly genome and discovered a new protein that defines their relationship to the DNA sequence.
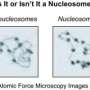
In organisms that store their genomic DNA in a nucleus, the genetic information is packaged into a condensed form called chromatin. In chromatin, the DNA is wound around particles called nucleosomes, each made up of four histone proteins, and the resulting "spools" are separated from one another by linker DNA. Each of the cell's chromosomes can therefore be thought of as an open necklace in which the DNA forms the string and the nucleosomes are the pearls. The precise positioning of the nucleosomes with respect to the nucleotide sequence of each DNA molecule, and the distances between individual nucleosomes, can have important consequences for gene regulation, and must therefore be tightly controlled. Using the fruit fly Drosophila melanogaster as a model, a research team led by Professor Peter Becker at LMU's Biomedical Center has now systematically determined the arrangement of the nucleosomes along the DNA, and identified a novel protein that plays a role in specifying their positions. The findings appear in the new issue of the journal Molecular Cell.
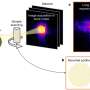
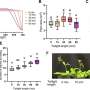
Previous investigations had shown that the nucleosomes at the start of active genes are often organized in a very orderly fashion, with a fixed length of DNA separating each one from the next. Such a non-random arrangement is known as a phased nucleosome array, and the phenomenon as such is referred to as nucleosome phasing. "These groups of nucleosomes could be ordered in such a way as to leave important gene regulatory sequences in the DNA exposed," explains Sandro Baldi, first author of the study. "So we searched specifically for these arrays in the genome of Drosophila." This essentially involved the assembly of a genome-wide map of the positions of all the nucleosomes in the fruit fly. With the aid of newly developed ways of analyzing the data, Becker and his team were able to identify and catalog all the sites at which phased nucleosomal arrays are found.
It turns out that about half of the regularly disposed nucleosomes were located close to active genes. This result confirmed that phasing does have something to do with gene activity. "But to our surprise," says Baldi, "the other half of the regularly ordered nucleosome sets were not associated either with gene sequences or with other known regulatory DNA regions."
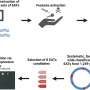
To learn more about the mechanisms that underlie nucleosome positioning, Becker and his colleagues developed a new method for the assembly of Drosophila chromatin in the test tube. They first isolated fruit fly DNA, free of proteins, and then added a protein extract obtained from Drosophila eggs. This extract contains all the factors required for the development of the early embryo, including histones and non-histone proteins that bind to chromatin.
"Within an hour of adding the extract to purified genomic DNA, the constituents of the mixture autonomously formed essentially the same chromatin structure as we find in Drosophila cell nuclei," says Becker. In particular, the researchers confirmed that the phased nucleosome groups which are not associated with active genes are correctly placed during the self-assembly process. And in further experiments, they came across a previously uncharacterized protein that binds directly to the naked DNA and triggers the formation of a regularly spaced group of nucleosomes around it.
"This new protein presumably plays an important role in genome organization, since it is one of the major factors responsible for phasing, which is why we named it Phaser," says Becker. "It is not yet clear whether mammals possess a similar protein. At all events, it will be very interesting to elucidate the functions of such phaser proteins. Our method for in-vitro reconstitution of chromatin will enable us to obtain insights into other mechanisms of genome organization, and can be used for further detailed studies of many other aspects of genome biology."
More information: Sandro Baldi et al, Genome-wide Rules of Nucleosome Phasing in Drosophila, Molecular Cell (2018). DOI: 10.1016/j.molcel.2018.09.032
Journal information: Molecular Cell
Provided by Ludwig Maximilian University of Munich