Molecular biologists compared human and yeast FACT
A protein complex called facilitates chromatin transcription (FACT) plays a role in DNA packing within a nucleus, as well as in oncogenesis. A team of scientists from MSU, working in cooperation with foreign colleagues, have reported similarities between the work of this complex in humans and yeast. This discovery led to the prediction that a new protein assists the FACT complex in humans. An article about the study was published in Journal of Biological Chemistry.
In eukaryotes, hereditary information is encoded in DNA, which is about one meter long, but is packed within the nucleus. The DNA molecule is extremely thin, and if it were chaotically crumpled, it would be impossible to detangle it without damage. In order to read the genetic code, it has to be unpacked. Histones, the proteins that bind to DNA, help to pack the genome in the cell nuclei. This complex is called chromatin. A minimal unit of chromatin is called a nucleosome; chromatin looks like a sequence of sewing spools (nucleosomes) with two DNA loops tightly wrapped on each of them.
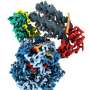
However, the work of this system also includes lots of other proteins. One of the protein complexes that control this process is FACT . It secures more effective RNA reading from the DNA template packed in chromatin.
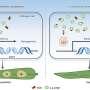
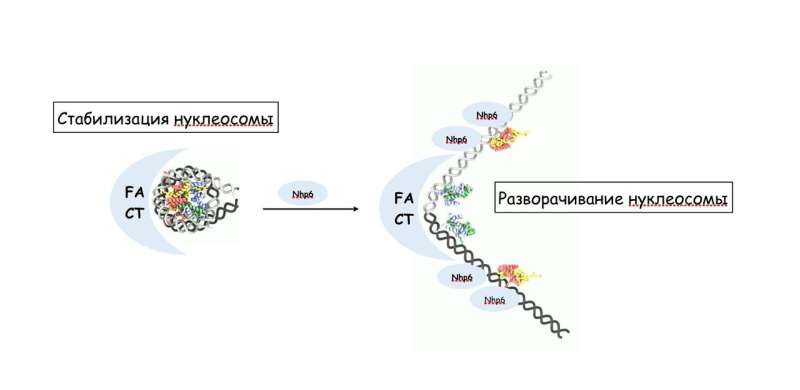
"On the molecular level, yeast FACT can both stabilize nucleosomes in the genome during RNA reading and unfold nucleosomes," said study co-author Vasily Suditsky, the head of the laboratory for the regulation of transcription and replication of the Faculty of Biology, MSU. "The balance between these two functions is determined by the presence of the yeast protein Nhp6, which helps FACT unfold nucleotides. Until recently, the second function was unknown for human FACT, despite the considerable similarity of the structure of these protein complexes in yeast and humans."
Surprisingly, FACT straightens the DNA looped around the spools without expending energy. Previously, such studies were conducted using yeast material. In the new article, Vasily Suditsky and his colleagues analyzed the mechanism of operation of this complex in humans. They were the first to prove that the Nhp6 protein from yeast in combination with human FACT complex is able to unfold nucleosomes.
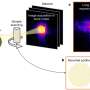
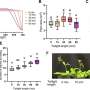
"The critical data were obtained at the Faculty of Biology, MSU using a modern biophysical approach—single-particle FRET (spFRET is a method that allows the measurement of the interatomic distance within one DNA-protein complex," said Studitsky. "The information we received led to the assumption that humans may possess a homologue of the yeast protein Nhp6 helping FACT unfold nucleosomes and regulate gene transcription."
The team also found important differences between human and yeast FACT complexes, though the activity of both varieties appeared to be relatively similar. The scientists found that in vitro yeast FACT cannot work without Nhp6, but in vivo (in cells), it functions. However, in this case, the functions of the FACT complex may be compensated with other proteins. Therefore, it is still unknown whether yeast FACT can function without Nhp6. On the contrary, human FACT is not stably associated with nucleosomes without additional proteins that haven't been studied in humans yet.
FACT complex is evolutionarily conserved. Scientists tried adding yeast Nhp6 to human FACT complex, and it functioned. Therefore, it can be assumed that humans also have a still unknown protein that helps FACT function.
"In the future, we plan to search for this hypothetical Nhp6-like human protein and to determine the role of nucleosome unfolding by FACT in gene regulation and detection of regions with damaged chromatin structure in the genome," said Studitsky.
More information: Laura L. McCullough et al, Functional roles of the DNA-binding HMGB domain in the histone chaperone FACT in nucleosome reorganization, Journal of Biological Chemistry (2018). DOI: 10.1074/jbc.RA117.000199
Journal information: Journal of Biological Chemistry
Provided by Lomonosov Moscow State University