Fujitsu develops molecular simulation technology to effectively create new drug candidates
Fujitsu Laboratories today announced the development of molecular simulation technology for drug discovery that can accurately estimate binding affinity, which represents the degree to which proteins that can cause diseases (target proteins) bind to chemical substances that could become candidate drugs. In the process of drug discovery, there is a demand for accurate prediction of the binding affinity between target proteins and chemical substances, which offers a rough estimate of a drug's efficacy. Molecular simulation technology has been widely used in the past as a method of predicting binding affinity, calculating the approximate forces that arise between atoms in molecules using Newtonian mechanics. The problem with this method, however, remains that the low degree accuracy of its estimation of the most important parameters-the degree of torsion at the binding sites. This means that the accuracy of its estimation of the overall binding affinity is also poor.
Now, Fujitsu Laboratories has developed molecular simulation technology that estimates the degree of torsion in a chemical substance, which is directly connected to the predicted binding affinity. The new technology not only takes into account the bonding location where the torsion will occur, but also the impact of neighboring atoms. Fujitsu Laboratories evaluated this technology for 190 types of chemical substances, comparing the results with correct results arrived at from first principles calculation and then evaluating the error rate. Upon doing so, it was able to confirm that the error rate in the estimate of the degree of torsion was, on average, one-tenth that of previous technology. It is anticipated that the use of this new technology in IT-based drug discovery, with its ability to accurately estimate the binding affinity of targeted proteins and chemical substances, offers the potential for groundbreaking new drug discovery efforts that could not be achieved with previous approaches.
The discovery of new drugs requires significant expenses and timeframes that can be measured in decades, leading to a global search for new methods of discovering drugs. One of the methods that has received considerable interest is IT-based drug discovery, a new drug discovery method using computers that makes it possible to create chemical substances as candidates for novel drugs with a high probability of success. IT-based drug discovery has become a focal point for expectations as a groundbreaking technology for the creation of new drugs, because unlike previous methods of trial and error, in which chemical substances are repeatedly created and tested, this approach makes it possible to virtually design chemical substances and estimate their effects.
The effects of a chemical substance as a drug are expressed when the chemical substance binds to a target protein. When the chemical substance binds to the target protein, it can change its shape in line with that of the target protein. The degree of deformation, namely, the parameters that indicate the extent of this shape change, is directly connected to the binding affinity of the substance and the protein, and gives a rough idea of its effect as a drug. Given this, there is a strong demand for the ability to accurately predict this value. To calculate the degree of deformation of a chemical substance, there are methods based on quantum mechanics and methods based on Newtonian mechanics. Quantum-mechanics based first principles calculation enables extremely accurate calculations, resolving the states of electrons from the types and positions of the atoms involved. On the other hand, however, the ability of first principles to perform exacting calculations necessarily leads to massive time required to complete the calculations. In order to simulate the degree of deformation for numerous chemical substances, time required is in the order of years, making this method impractical. On the other hand, approximate calculations based on molecular simulations are extremely fast, using Newtonian mechanics to calculate the forces between the atoms within the molecules, and can even handle large molecules like proteins quite easily. Consequently, this method is widely used. With Newtonian mechanics, the forces between the atoms are expressed in the following manner:
- As a force that depends on the distance between two atoms bonded to each other
- A force that relies on the angles between three atoms bonded to each other
- A force that relies on the degree of torsion in the bond, and
- A force that relies on the distance between atoms that are not bonded.
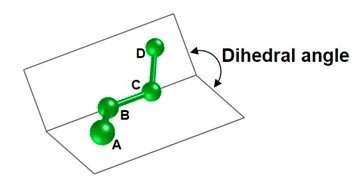
Among these, when a chemical substance is bound to a target protein, the degree of torsion of the bond represents the important degree of deformation. With existing technology, however, the accuracy of the estimation of the dihedral angle (figure 1) parameter, which is necessary to calculate the degree of torsion of the bond, is quite low, resulting in the problem of low accuracy in the estimation of the affinity of the bond in the simulation.
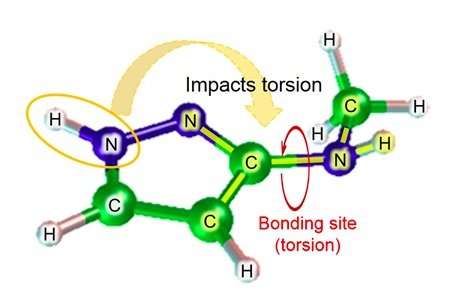
Fujitsu Laboratories has been developing molecular simulation technology for more than ten years. Now, using the knowledge it obtained through previous efforts, Fujitsu Laboratories has developed a molecular simulation technology that can estimate the dihedral angle parameter by taking into consideration the impact of atoms near the bond. Existing technology estimates the dihedral angle parameter based on a total of four atoms-the two atoms in the relevant bond, and the other atoms each of those atoms was bonded to. Depending on the structure of the molecule, however, there are cases where atoms beyond those four could have a significant impact, and in those cases, the margin of error of the estimation could be quite large. With this technology, Fujitsu Laboratories has created a database of estimation formulas for partial structure patterns where the impact of atoms further away from the bond site could be significant, as well as for the degree of torsion of chemical substances that would be expected in that case. Using the relevant estimation formula to find the degree of torsion (figure 2) in the case of molecules corresponding to the database for partial structures, it has become possible to even make highly accurate estimations for molecular torsion, which was previously difficult to calculate accurately.
When Fujitsu Laboratories integrated this technology into the software it had developed for generating sophisticated parameters for the forces between atoms (FF-FOM), it was able to confirm that the results conformed to accurate computations.
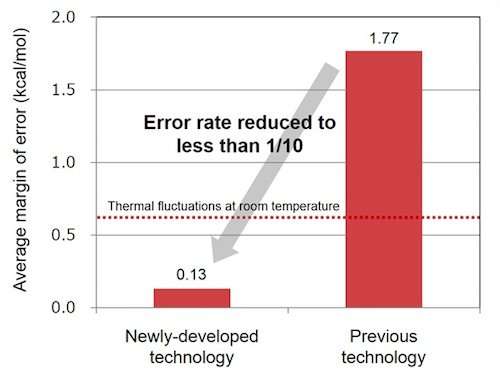
When Fujitsu Laboratories evaluated the difference between the results of this technology and the results of a calculation from first principles for the estimation of the degree of torsion with 190 types of chemical substances, it was less than one-tenth that of the previous technology, on average, 0.6 kcal/mol below room temperature thermal fluctuations, confirming that the new technology is practical. Because it can accurately estimate the binding affinity of target proteins and chemical substances, it is expected that the use of this technology will lead to the creation of groundbreaking new drugs through its use in IT-based drug discovery.
Provided by Fujitsu