Computer taught to intuitively predict chemical properties of molecules
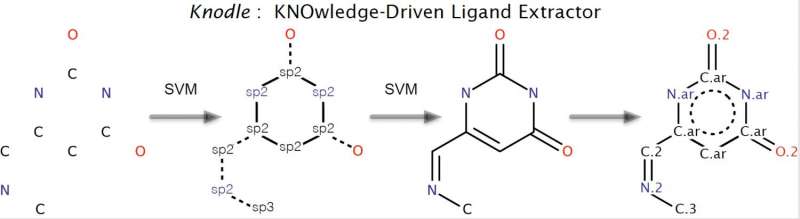
Scientists from MIPT's Research Center for Molecular Mechanisms of Aging and Age-Related Diseases, together with Inria research center, Grenoble, France, have developed a software package called Knodle to determine an atom's hybridization, bond orders and functional groups' annotation in molecules. The program streamlines one of the stages of developing new drugs. A paper on the new development has been published in the Journal of Chemical Information and Modeling.
Once a drug has entered the human body, it needs to act on the cause of a disease. On a molecular level, disease is a malfunction of some proteins and their encoding genes. In drug design, these are called targets. If a drug is antiviral, it must somehow prevent the incorporation of viral DNA into human DNA. In this case, the target is a viral protein. The structure of the incorporating protein is known as the active site. Inserting a molecular "plug" prevents the viral protein from incorporating itself into the human genome and the virus will die. Thus, you find the "plug," you have your drug.
To find the required molecules, researchers use an enormous database of substances. There are special programs capable of finding a needle in a haystack; they use quantum chemistry approximations to predict the place and force of attraction between a molecular "plug" and a protein. However, databases only store the shape of a substance; information about atom and bond states is also needed for an accurate prediction. Determining these states is what Knodle does. With the help of the new technology, the search area can be reduced from hundreds of thousands to just 100. These 100 can then be tested to find drugs such as Reltagravir, which has been used for HIV prevention since 2011.
Students are used to seeing organic substances represented as letters with sticks (substance structure), though in reality, there are no sticks. Every stick is actually a bond between electrons that obeys the laws of quantum chemistry. In the case of one simple molecule like the one in the diagram, the experienced chemist intuitively knows the hybridizations of every atom (the number of neighboring atoms it is connected to) and after a few hours looking at reference books, she can reestablish all the bonds. This is because hundred of similar substances are known, and if oxygen is "sticking out like this," it almost certainly has a double bond. In their research, Maria Kadukova, an MIPT student, and Sergei Grudinin, a researcher from Inria research center located in Grenoble, France, decided to pass on this intuition to a computer by using machine learning.
Compare the phrases "A solid hollow object with a handle, opening at the top and an elongation at the side, at the end of which there is another opening," and "a vessel for the preparation of tea." Both of them describe a teapot, but the latter is simpler. The same is true for machine learning—the best algorithm for learning is the simplest. This is why the researchers chose to use nonlinear support vector machines (SVM), a method used in recognizing handwritten text and images.
Machine learning requires a lot of examples, and the scientists did this using 7605 substances with known structures and atom states. "This is the key advantage of the program we have developed— learning from a larger database gives better predictions. Knodle is now one step ahead of similar programs. It has a margin of error of 3.9 percent, while for the closest competitor this figure is 4.7 percent," explains Maria Kadukova. And that is not the only benefit. The software package can easily be modified for a specific problem. For example, Knodle does not currently work with substances containing metals, because those kind of substances are rather rare. But if it turns out that a drug for Alzheimer's is more effective with a metal, the only thing needed to adapt the program is a database with metallic substances.
More information: Maria Kadukova et al, Knodle: A Support Vector Machines-Based Automatic Perception of Organic Molecules from 3D Coordinates, Journal of Chemical Information and Modeling (2016). DOI: 10.1021/acs.jcim.5b00512
Journal information: Journal of Chemical Information and Modeling
Provided by Moscow Institute of Physics and Technology