Researchers observe the switching of Ras protein in detail

Ras proteins are molecular switches that decide if and when cells divide inside our bodies. An impairment of their function may result in the formation of a tumour. The process of switching the proteins on and off has been observed in detail by a research team headed by Prof Dr. Klaus Gerwert from the Department of Biophysics at Ruhr-Universität Bochum (RUB); using a combination of methods, the team confirmed the hypothesis that the binding partner of Ras in its bonded form does not contain any hydrogen atoms in the phosphate groups. The renowned Journal of Biological Chemistry has published the report as its cover story on March 16, 2018.
Possible causes of cancer
Ras proteins function as small switches: depending on whether they are bonded with guanosine triphosphate (GTP) or with guanosine diphosphate (GDP), they are either switched on or off. The GTP bond triggers complex signalling pathways that lead to the nucleus where they initiate cell division. Subsequently, Ras proteins are deactivated by cleaving off a phosphate group from the bonded guanosine triphosphate molecule and generating guanosine diphosphate.
If this process is inhibited, for example due to mutations in the Ras protein, Ras remains in its active state, and the ongoing division of cells can lead to the formation of a tumour. "More than 30 per cent of all tumours carry a mutation of the Ras protein," explains Dr. Daniel Mann from the research team. "Consequently, the cleavage reaction of GTP in Ras is the key for understanding many types of cancer."
How many hydrogen atoms?
In order to understand GTP cleavage reactions in Ras, the researchers need a precise image of the starting point, i.e. they must know what Ras-bound GTP looks like. This includes the question if the three phosphate groups of the GTP molecule contain any hydrogen atoms, since they function as acids and, for example, carry a hydrogen atom when dissolved in water.
However, it is for the most part not possible to measure hydrogen directly using conventional methods such as X-ray crystallography. There is a considerable difference between the chemical reactions of GTP cleavage with or without bonded hydrogen. It has been generally assumed that the phosphate groups of GTP-bound Ras are free of hydrogen atoms. This assumption is however based not on direct measurements, but on the interpretation of changes in the measured infrared and nuclear magnetic resonance spectrums.
Common assumption doubted
Recent neutron diffraction analyses of the GTP-analogue GppNHp have triggered doubts in this universally accepted hypothesis. In the process, evidence has emerged that GTP might contain hydrogen atoms after all. "This has challenged all notions of GTP cleavage to date," says assistant professor Dr. Carsten Kötting.
3-D films with subatomic resolution
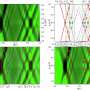
The RUB researchers have performed further measurements, in order to record both the infrared spectrums of the GppNHp that is deployed in neutron diffraction and the spectrums of the natural GTP in Ras. Thus, it was possible to draw a direct comparison with neutron diffraction, as well as a comparison with an environment that resembles that of the human cell. "The measured infrared data enable us to decode molecular reactions at the highest temporal and spatial resolution," elaborates Klaus Gerwert. "However, the information is coded in infrared spectra and is therefore difficult to interpret."
The RUB researchers deploy a process for decoding that facilitates the calculation of experimental data such as infrared spectra and nuclear spin spectra gathered through coupled quantum mechanics/molecular mechanics simulations. "If the calculated spectra correspond with the spectra measured in experiments, we can translate the figures we measured in experiments into three-dimensional films," says Daniel Mann.
Hypothesis confirmed
The films have a resolution smaller than a tenth of the atomic diameter. "These precise results have enabled us to finally determine the accurate protonation state of GTP bound to Ras proteins: the GTP phosphate groups do indeed not carry any hydrogen atoms," concludes Klaus Gerwert.
More information: Daniel Mann et al. The protonation states of GTP and GppNHp in Ras proteins, Journal of Biological Chemistry (2018). DOI: 10.1074/jbc.RA117.001110
Journal information: Journal of Biological Chemistry
Provided by Ruhr-Universitaet-Bochum