Simple method measures how long bacteria can wait out antibiotics
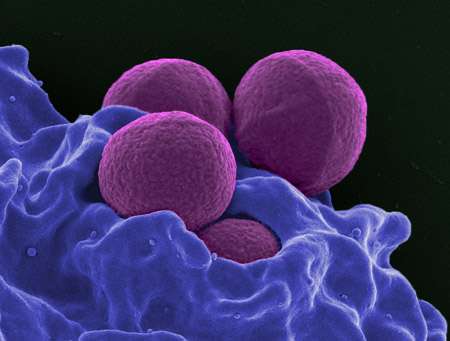
A growing number of pathogens are developing resistance to one or more antibiotics, threatening our ability to treat infectious diseases. According to a study published June 20th in Biophysical Journal, a simple new method for measuring the time it takes to kill a bacterial population could improve the ability of clinicians to effectively treat antimicrobial-tolerant strains that are on the path to becoming resistant.
"These findings allow measurement of tolerance, which has previously been largely overlooked in the clinical setting," says senior study author Nathalie Balaban of The Hebrew University of Jerusalem. "Routinely measuring tolerance could supply valuable information about the duration of antibiotic treatments, reducing the chance of both under- and over-treatment. Furthermore, data compiled from such measurements could give an estimate of how widespread the phenomenon of tolerance really is, which is currently a complete unknown."
According to the World Health Organization, antibiotic resistance is one of the biggest threats to global health and is putting the achievements of modern medicine at risk. Due to selective pressure, pathogens acquire resistance through mutations that make the antibiotic less effective, for example, by interfering with the ability of a drug to bind to its target. Currently, clinicians determine which antibiotic and dose to prescribe by assessing resistance levels using a routine metric called minimum inhibitory concentration (MIC)—the minimal drug concentration required to prevent bacterial growth.
Although resistant strains continue to grow despite exposure to high drug concentrations, tolerant strains can survive lethal concentrations of an antibiotic for a long period of time before succumbing to its effects. Tolerance is often associated with treatment failure and relapse, and it is considered a stepping stone toward the evolution of antibiotic resistance. But unlike resistance, tolerance is poorly understood and is currently not evaluated in healthcare settings.
"The lack of a quantitative measure means that this aspect of the treatment relies largely on the experience of the individual physician or the community," says first author Asher Brauner, a PhD student in Balaban's lab. "This can lead to treatment being either too short, increasing the risk of relapse and evolution of resistance, or much too long, unnecessarily causing side effects, release of antibiotic waste into the environment, and additional costs."
To address this problem, Balaban and her team developed a tolerance metric called the minimum duration for killing 99% of the population (MDK99). The protocol, which can be performed manually or using an automated robotic system, involves exposing populations of approximately 100 bacteria in separate microwell plates to different concentrations of antibiotics for varied time periods, while determining the presence or lack of survivors.
The researchers applied MDK99 to six Escherichia coli strains, which showed tolerance levels ranging from 2 to 23 hr under ampicillin treatment. MDK99 also facilitates measurements of a special case of tolerance known as time-dependent persistence—the presence of transiently dormant subpopulations of bacteria that are killed more slowly than the majority of the fast-growing population. Like other forms of tolerance, time-dependent persistence can lead to recurrent infections because the few surviving bacteria can quickly grow to replenish the entire population once antibiotic treatment stops.
"A take-home message from this is that it is important to complete a course of antibiotic treatment as prescribed, even after the disappearance of the symptoms," Balaban says. "Partial treatment gives tolerance and persistence mutations a selective advantage, and these, in turn, hasten the development of resistance."
In future studies, Balaban and her team will use MDK99 to study the evolution of tolerance in patients. Moreover, the ability to systematically determine the tolerance level of strains in the lab could facilitate research in the field. "If implemented in hospital clinical microbiology labs, MDK99 could enable the efficient classification of bacterial strains as tolerant, resistant, or persistent, helping to guide treatment decisions," Balaban says. "In the end, understanding tolerance and finding a way to combat it could significantly reduce the ever-growing risk of resistance."
More information: Biophysical Journal, Brauner et al.: "An Experimental Framework for Quantifying Bacterial Tolerance" www.cell.com/biophysj/fulltext … 0006-3495(17)30551-9 , DOI: 10.1016/j.bpj.2017.05.014
Journal information: Biophysical Journal
Provided by Cell Press