Plastic proteins: Synthetic material mimics essential characteristics of natural proteins
Researchers hoping to design new materials for energy uses have developed a system to make synthetic polymers—some would say plastics—with the versatility of nature's own polymers, the ubiquitous proteins. Based on an inexpensive industrial chemical, these synthetic polymers might one day be used to create materials with functions as limitless as proteins, which are involved in every facet of life.
Reporting in Angewandte Chemie International Edition March 14, researchers reveal a method to produce polymers that mimic proteins in the versatility of their raw ingredients and how those ingredients link together to form a larger structure.
"Proteins are sequence-defined polymers and have a whole variety of exquisite functions," said materials scientist Jay Grate of the Department of Energy's Pacific Northwest National Laboratory. "But natural materials are unstable. That's good for nature, but if we want stable, long-lasting materials, we need to make our own sequence-defined polymers."
Stuff of life
Proteins are at the core of life: In living things, they are architect and engineer. They are the wrenches and machines that build an organism's varied parts, building those parts out of other proteins of many sizes and shapes. They form the power plants in cells, run the plants, make energy and store energy. They make things grow, and are the bricks of growth as well.
Because of their versatility, proteins are some of researchers' favorite tools. Many drugs are re-engineered proteins such as converted antibodies (for example, drugs whose names end in -mab). The problem, however, is that proteins are also short-lived. Nature designed them to be temporary and recyclable. Any environment proteins find themselves in is full of things—often other proteins—that break them down.
One way to get around these butchers is to design a material that behaves like proteins but is not actually protein. To that end, researchers are pursuing materials that mimic the building blocks of proteins—amino acids. Amino acids bestow on proteins their tremendous variety and versatility.
Those qualities are what plastic protein researchers are trying to emulate. Amino acids come in 20 or so variations. Each has the same backbone, from which juts a group of atoms called a side chain that gives the amino acid its particular chemical characteristic. The amino acid backbones snap together like beads on a string, the side chains arranged in a particular order for each protein.
But proteins aren't floppy pearl necklaces. The beads fold in upon themselves to form structured objects. Some proteins end up looking like balls, some like capital Ys, others like olive wreaths.
These shapes come about because the side chains and the protein backbone stick to other side chains and backbone regions like Post-Its stick to one another. The folding and sticking are very specific, like origami, resulting in a particular structure rather than a tangled mess.
Threes
Grate needed three things to mimic proteins: raw components with a backbone that can support a large variety of side chains; the ability to put the side chains in a particular order; the stickiness, which chemists refer to as non-covalent bonds.
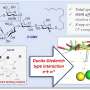
He had been working with an industrial chemical called cyanuric chloride for unrelated purposes, but his understanding of its chemical properties made him think it might be a good starting point. Cyanuric chloride is a molecule that has three convenient places it could be extended. Two of them can link together, like two people holding hands, to form the backbone. The third can house a side chain. All together, Grate called the resulting molecule a triazine-based polymer, or TZP for short.
Although such a polymer would be immune to protein-destroying entities, Grate expects other things in the environment such as bacteria will break it down, based on TZP's chemical nature. So the material would last, but not forever.
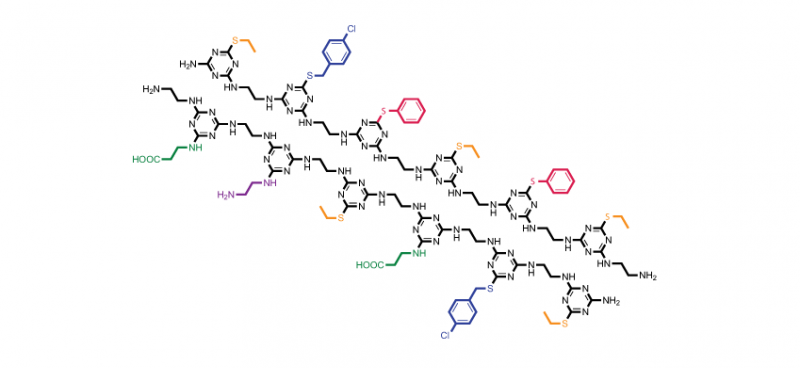
Idea in hand, Grate and PNNL chemist Kai-For Mo had to develop a way to synthesize TZP. They made a variety of monomers by adding different side chains to cyanuric chloride, with each monomer a single building block analogous to an amino acid. For this study, they created five different side chains. Then they found they could add one monomer at a time relatively simply by changing the temperature at which they performed the chemical reactions, among other synthesis tricks.
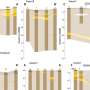
After synthesizing polymers six monomers long, called a 6-mer in polymer parlance, the researchers verified their creations. They used analytical instruments to show that the polymers were the right size, had the right side chains, and the side chains were in the right order. They also synthesized a 12-mer to show this method works with longer polymers.
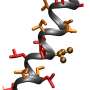
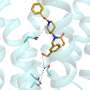
To see whether TZPs would fold in a manner analogous to proteins, PNNL computer scientist Michael Daily simulated small TZPs, singly and interacting with each other. A 6-mer folded neatly in half, forming a straight rod three monomers long and two monomers wide, like a hairpin. Similarly, two 3-mers lined up along each other, partially intertwined like a zipper.
The stickiness holding these "nanorods" together were non-covalent bonds between backbone atoms, the same types of bonds nature uses so that proteins take their proper shapes. And like protein structures, the TZP side chains were arranged in specific positions around the exterior of the rods formed by the backbone-backbone interactions.
The next step is to create a larger library of side chains, the count of which is ten so far. Then they must make longer polymers and show that they really do take useful shapes. Once researchers understand the rules for how to get specific shapes with TZPs that also assemble into larger structures, they can design materials with desired functions—for example, a membrane for a battery, a catalyst for a fuel cell, or even a therapeutic drug.
More information: Jay W. Grate, Kai-For Mo, and Michael D. Daily. Triazine-based Sequence Defined Polymers with Side Chain Diversity and Backbone-Backbone Interaction Motifs, Angewandte Chemie International Edition, March 14, 2016, DOI: 10.1002/anie.201509864
Journal information: Angewandte Chemie International Edition
Provided by Pacific Northwest National Laboratory