Team tackles mystery of protein folding
Proteins are the workhorses of life, mediating almost all biological events in every life form. Scientists know how proteins are structured, but folding - how they are built - still holds many mysteries.
New research conducted at Michigan State University and published in the current issue of Nature Chemical Biology, features a chemistry approach that's solving some of the riddles of the complex protein-building process of folding. When it goes right, strings of amino acids become well-ordered, three-dimensional proteins in a split second. When it goes awry, though, it's the first step of many serious diseases.
When errors happen in folding, proteins clump together, form plaques such as those found in Parkinson's disease and cystic fibrosis, and cause cells to degenerate. Understanding folding could lead to medicinal advances to treat these and other diseases at their earliest stage.
"Our novel tool set can potentially be applied to analyze how disease mutations impact the structural and functional integrity of pathologically important membrane proteins," said Heedeok Hong, MSU chemist and study co-author. "This knowledge will ultimately help in designing treatments that can stabilize defective membrane proteins for their optimal function."
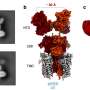
The team focused on membrane proteins because roughly 30 percent of all proteins reside in this oily layer that encapsulates cells. Membrane proteins carry out many life functions, including the uptake of nutrients, secretion of wastes, maintaining ion balance and transmitting nerve signals.
"Despite their importance, we know little about how membrane proteins fold because studying membrane protein folding has been formidable, due to the lack of adequate methods," Hong said.
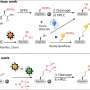
To tackle the membrane mysteries, the team developed a new method called "steric trapping." First, scientists attached two small molecular tags to a protein in its folded form. Next, they added bulky objects that bind the tags. The large attachments, by their sheer size alone, unravel the protein to its unfolded state.
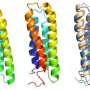
This simple yet eloquent procedure can test the stability of membrane proteins, show what unfolded membrane proteins look like and reveal how individual amino acids that are building a protein work together to maintain its folded shape.
"Using this novel tag-binding system, or steric trapping, our team was able to observe and test membrane proteins without disturbing their native environment," Hong said. "Controlling folding and unfolding while keeping their native membrane environment has been one of the major methodological hurdles to solve the membrane protein folding problem. We have overcome one, and now we are ready for another."
More information: Ruiqiong Guo et al. Steric trapping reveals a cooperativity network in the intramembrane protease GlpG, Nature Chemical Biology (2016). DOI: 10.1038/nchembio.2048
Journal information: Nature Chemical Biology
Provided by Michigan State University