Improved methods provide insight into the nature of substrate, ligand and lipid interactions
Membrane proteins account for up to 30% of the proteins present in an organism, though relatively few are well characterised compared to their soluble protein counterparts. This is largely due to their instability when extracted from membranes using detergent, which hampers their study. Methods are required to assess the integrity of extracted protein, which are not always straightforward or reliable.
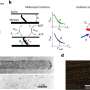
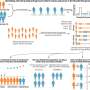
Researchers, led by Dr Edmund Kunji at the Mitochondrial Biology Unit in Cambridge, have adapted a fluorescent dye-based method of assessing membrane protein stability for rapid and high throughput use. The new assay requires relatively little sample and uses low specification equipment that is suitable for use with detergents, providing significant advantages for anyone working with membrane proteins.
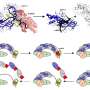
The team report that as well as highlighting optimal stabilising conditions, the detailed trends in stability gained using the technique reveal insights into membrane protein interactions with substrate, lipids, and state-specific inhibitors. When applied to the ADP/ATP carrier, a small membrane protein that plays a fundamental role in energy metabolism, the group could demonstrate that particular lipids interact with the protein by two distinct mechanisms, and reveal a previously unconfirmed interaction of lipid in a particular state of the protein.
The technique also provided clues on how Uncoupling protein 1, a protein that may help combat obesity, operates in mitochondria, which is currently debated. Importantly, the assay provides a quick way to discern folded protein from unfolded protein, which is particularly useful in membrane protein research. As a rapid high throughput technique, the researchers believe that the procedure will have wider applications too, for example to identify the function of membrane proteins that are currently unknown. Investigations into the versatility of the method are on going.
More information: "Trends in thermostability provide information on the nature of substrate, inhibitor, and lipid interactions with mitochondrial carriers." J Biol Chem. 2015 Mar 27;290(13):8206-17. DOI: 10.1074/jbc.M114.616607
Provided by MRC Mitochondrial Biology Unit