July 24, 2015 report
Why do mitochondria retain their own genome?
It sounds like science fiction to suggest that every cell in the human body is occupied by a tiny genome-equipped organelle, with which we exist in symbiosis. But in actuality, eukaryotic life is dependent on mitochondria, which provide energy to the cell in the form of adenosine triphosphate (ATP). Over millennia, the genomes of mitochondria evolved under selection for minimal gene content, but researchers have been unable to determine why some but not all mitochondrial genes have been transferred to the nuclear genome.
An international collaborative of researchers formed an interesting hypothesis regarding this phenomenon: The mitochondrial genome encodes hydrophobic membrane proteins which, if encoded in the nucleus, would be filtered by the signal recognition particle (SRP) and misdirected into the endoplasmic reticulum. The researchers conducted a study exploring their hypothesis and have published their results in the Proceedings of the National Academy of Sciences.
In order to predict if a protein would be targeted by SRP, the researchers calculated the free insertion energy of transmembrane proteins, which, if scored zero or less, were considered to be hydrophobic. Higher scores were rated marginally hydrophobic. If a transmembrane protein domain (TMD) scored hydrophobic and the length of its tail was longer than 120 amino acids, the researchers predicted it would be arrested by SRP and directed into the endoplasmic reticulum.
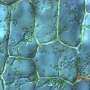
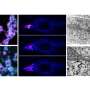
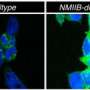
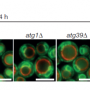
They expressed such proteins in cellular cytoplasm and were able to determine that they were, in fact, arrested by the SRP and directed to the endoplasmic reticulum. Further, the researchers observed that the mistargeting of these hydrophobic proteins into the soluble medium of the endoplasmic reticulum resulted in the formation of aberrant honeycomb structures similar to those observed during viral infections. "We conclude that genes for hydrophobic membrane proteins of more than 120 amino acids are likely retained in distinct organelle genomes to ensure a correct localization of these proteins and avoid transport to the endoplasmic reticulum," the authors write.
Thus, the researchers conclude, the selection against mistargeting hydrophobic proteins into the endoplasmic reticulum posed at least one major selective constraint on the retention of the mitochondrial genome. They bolster this finding by comparing it to similar membrane dynamics in the chloroplasts of plants.
Previous studies have suggested that one-third of mitochondrial proteins have evolved in response to the specific environmental constraints of different species. Most of these proteins are involved with transport, regulatory, and membrane functions. The results of the current study are consistent with these findings.
A persistent mystery has been the evidence that in rare cases, transfers of otherwise universal mitochondrial genes into the nuclear genome have occurred. The results of the current study present an explanation: These particular proteins demonstrate reduced hydrophobicity in those species in which the transfer took place.
Clinical applications include the development of methods to cure human mitochondrial genetic diseases, such as Leber's hereditary optic neuropathy. "These results resolve the long-standing question about why aerobic mitochondria and photosynthetic chloroplasts need a distinct compartmental genome, by and large, although other factors may be involved," the authors write.
More information: "Mitochondrial genomes are retained by selective constraints on protein targeting." PNAS 2015 ; published ahead of print July 20, 2015, DOI: 10.1073/pnas.1421372112
Journal information: Proceedings of the National Academy of Sciences
© 2015 Phys.org