Structural data reveals new mechanism behind protein transport
In order for newly-produced secretory or membrane proteins to find their final destination, the proteins have signal-sequences connected to themselves as a form of address tag. Furthermore, they use a particle guiding them to the cell membrane. In a new study, Umeå researchers show how this particle recognizes these address tags and thus makes it possible for the proteins to be directed to the cell membrane. The results are published in an article in the journal Nature Communications.
To function, most of the newly-produced proteins first need to be sorted and then transported within and outside of the cell. For this purpose, proteins have a signal-sequence bound to it that is recognised by other proteins guiding them to their final destination.
One of these protein transporters is the so-called signal recognition particle (SRP). Researchers at Umeå University show in a new, detailed structural study, published in the journal Nature Communications, how the binding of the signal-sequence triggers a number of structural changes within the SRP particle. These changes make it possible for the protein to be transported further to the cell membrane.
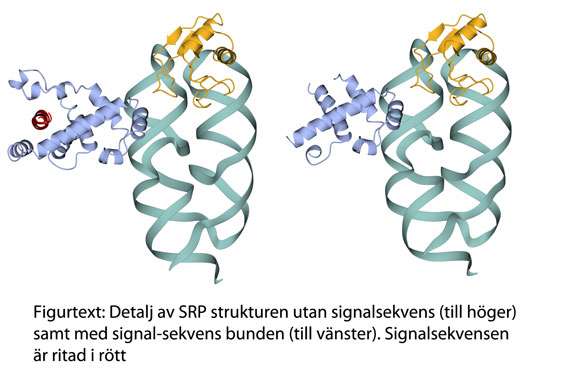
By studying and comparing the structure of unbound SRP with a signal-sequence bound SRP, a detailed picture of the impact of this binding is now available. The research team has in several previous studies, based on x-ray crystallographic methods, shown the basic structure of SRP. This study is the continuation of previous research and shows more details of structural changes between unbound and signal-sequence bound SRPs.
"We can now show how parts of the SRP normally flexible in the structure become folded forming a binding fold perfectly adapted to the signal-sequence," says Elisabeth Sauer-Eriksson, Professor at the Department of Chemistry.
The structure of the SRP signal-sequence complex is determined using x-ray crystallographic methods. Using this method, the biological molecules' three-dimensional structure can be determined at a nearly atomic resolution.
The signal-sequences are highly diverse in both length and in amino-acid composition but they all have a core of at least eight consecutive hydrophobic amino acids that acts as the major determinant for recognition by SRP. Researchers are now continuing to clarify the details underlying the targeting mechanism. A yet to be answered question is: how it is possible for SRP to specifically bind to so many differing kinds of signal-sequences?
About SRP:
SRP is a ribonucleoprotein complex consisting of RNA and several protein components. SRP is present and conserved in all organisms; evidence for its fundamental role in all living cells.
More information: "Signal-sequence induced conformational changes in the signal recognition particle." Nature Communications 6, Article number: 7163 DOI: 10.1038/ncomms8163
Journal information: Nature Communications
Provided by Umea University


















