Researchers piece together how proteins fold

(Phys.org) -- A new method for looking at how proteins fold inside mammal cells could one day lead to better flu vaccines, among other practical applications, say Cornell researchers.
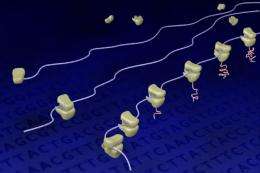
The method, described online in the Proceedings of the National Academy of Sciences July 16, allows researchers to take snapshots of the cell's protein-making machinery -- called ribosomes -- in various stages of protein production. The scientists then pieced together the snapshots to reconstruct how proteins fold during their synthesis.
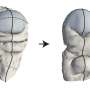
Proteins are made up of long chains of amino acids called polypeptides, and folding gives each protein its characteristic structure, which determines its function. Though researchers have used synthetic and purified proteins to study protein folding, this study looks at proteins from their inception, providing a truer picture for how partially synthesized polypeptides can fold in cells.
Proteins fold so quickly -- in microseconds -- that it has been a longtime mystery just how polypeptide chains fold to create the protein's structure.
"The speed is very fast, so it's very hard to capture certain steps, but our approach can look at protein folding at the same time as it is being synthesized by the ribosomes," said Shu-Bing Qian, assistant professor of nutritional sciences and the corresponding author on the paper. Yan Han, a postdoctoral associate in Qian's lab, is the paper's first author.
In a nutshell, messenger RNA (mRNA) carries the coding information for proteins from the DNA to ribosomes, which translate those codes into chains of amino acids that make up proteins. Previously, other researchers had developed a technique to localize the exact position of the ribosomes on the mRNA. Qian and colleagues further advanced this technique to selectively enrich only a certain portion of the protein-making machinery, basically taking snapshots of different stages of the protein synthesis process.
"Like a magnifier, we enrich a small pool from the bigger ocean and then paint a picture from early to late stages of the process," Qian said.
In the paper, the researchers also describe applying this technique to better understanding a protein called hemagglutinin (HA), located on the surface of the influenza A virus; HA's structure (folding) allows it to infect the cell.
Flu vaccines are based on antibodies that recognize such proteins as HA. But viruses have high mutation rates to escape antibody detection. Often, flu vaccines lose their effectiveness because surface proteins on the virus mutate. HA, for example, has the highest mutation rate of the flu virus' surface proteins.
The researchers proved that their technique can identify how the folding process changes when HA mutates.
"If people know the folding picture of how a mutation changes, it will be helpful for designing a better vaccine," Qian said.
"Folding is a very fundamental issue in biology," Qian added. "It's been a long-term mystery how the cell achieves this folding successfully, with such speed and with such a great success rate."
Co-authors include researchers at the National Institute of Allergy and Infectious Diseases.
The research was funded by the National Institute of Allergy and Infectious Diseases Division of Intramural Research, National Institutes of Health Grant, Ellison Medical Foundation Grant and U.S. Department of Defense Exploration-Hypothesis Development Award.
Journal information: Proceedings of the National Academy of Sciences
Provided by Cornell University