University of Illinois biophysicists measure mechanism that determines fate of living cells
(Phys.org) —New tension gauge tether (TGT) laboratory method developed at the University of Illinois at Urbana-Champaign has broad applications for research into stem cells, cancer, infectious disease, and immunology.
Cells in the human body do not function in isolation. Living cells rely on communication with their environment—neighboring cells and the surrounding matrix—to activate a wide range of cellular functions, including reproduction of new cells, differentiation of stem cells into distinct cell types, cell adhesion, and migration of white blood cells to fight bodily infections. This cellular communication occurs on the molecular level and it is reciprocal: a cell receives cues from and also transmits function-activating cues to its neighbors.
The mechanics of this type of cellular interaction have been studied extensively: receptors extending through the cell membrane are activated when they form a bond to specific molecules. Now for the first time, University of Illinois biophysicists at the Center for the Physics of Living Cells and the Institute for Genomic Biology have measured the molecular force required to mechanically transmit function-regulating signals within a cell.
The new laboratory method, named the tension gauge tether (TGT) approach, developed by Taekjip Ha with postdoctoral researcher Xuefeng Wang, and reported in the May 24, 2013, issue of the journal Science, has made it possible to detect and measure the mechanics of the single-molecule interaction by which human cell receptors are activated. The researchers used integrin, a cell membrane receptor protein that is activated when it bonds to a ligand molecule.
In the TGT approach, Ha and Wang repurposed DNA strands, using them as tethers for ligand molecules, to test the tension required to activate cell adhesion through integrin. The integrin bonds to the tethered ligand, and adhesion is activated only if the DNA tether does not rupture.
Taking advantage of the geometric characteristics of DNA's double helix form, the researchers were able to tune the strands to rupture at discrete tension levels: by varying the attachment points along the DNA strands, the force required for rupture was either low (unzipping the helix), high (shearing the strands), or intermediary (combination of unzipping and shearing).
"If you went fishing and a fish broke your 30-lb fishing line but not the 40-lb one, you would know that its strength was in the range of 30-40 pounds," explained Wang. "Here we applied the same strategy to measure the molecular tension applied by cells (the fish). Mammalian cells apply a force to activate cell membrane proteins called integrins which mediate cell adhesion. We immobilized ligand molecules (the bait) on a surface through molecular tethers (the fishing line) with defined tension tolerances, tunable from 10 pico Newton (pN) to 60 pN). After integrin-ligand binding, cells apply a force on the bonds, and we compare this force to the molecular tether strength by observing cell adhesion status."
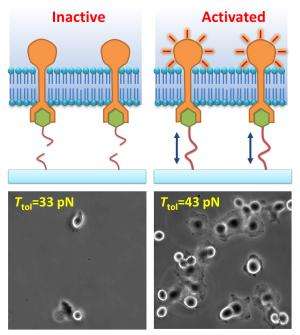
Since single-molecule interactions are difficult to monitor, the researchers observed the receptor-regulated cellular function, to gauge whether the integrin was activated. Ha and Wang discovered that integrin experiences a well-defined "quantum of force," about 40 pico-Newton (pN), to activate cell's adhesion to a surface.
"We observed that mammalian cells adhere on the culture surface with 43 pN tension tolerance of ligands, but not on 33 pN surface. Therefore we deduced that single molecular tension is around 40 pN on integrin cell-membrane receptors during cell adhesion," Wang added.
"This is a very exciting result," commented Ha. "With the ability to define the single molecular forces required to make living cells behave as desired, we may be one step closer to a remedy for certain hard-to-cure diseases. We know that the behavior of cancer cells and stem cells can be controlled by how stiff or soft their environments are. Understanding and manipulating molecular conversation through defined forces has huge implications for the development of future medical interventions. We expect the TGT approach will have broad applications in laboratory studies of cell differentiation, cancer metastasis, as well as immunology and infectious disease."
Journal information: Science
Provided by University of Illinois at Urbana-Champaign

















