New details about gene regulation explained
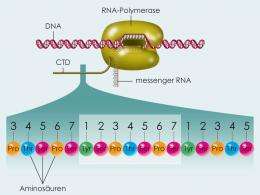
(Phys.org) -- When genetic information is read from the genetic blueprint DNA, RNA polymerase II translates it into RNA molecules. The C-terminal domain, abbreviated as CTD, is an important area of the polymerase molecule. It conveys information to the polymerase about how the genetic code should be copied and processed. To do this, the enzyme P-TEFb attaches molecular messages in the form of phosphate residues to certain positions within the CTD. Scientists at the Max Planck Institute of Molecular Physiology in Dortmund have described the pattern, on the basis of which P-TEFb completes this process. Their findings are not only important in terms of providing a better understanding of gene regulation in healthy organisms, they are also of medical interest: for example, HIV exploits this enzyme for its own purposes to accelerate the production of new virus proteins.
Despite their considerably more complex make-up, human beings do not have many more genes than a fruit fly, namely around 25,000. However, information is conveyed not only through the base sequence of the DNA, but also epigenetically in the form of small molecular attachments. For example, “molecular post-its” in the form of phosphate attachments tell the cell’s translation machinery which sections of the DNA should be read. Special enzymes that place such marks in the correct position play a crucial role in ensuring that the cell interprets the genetic code correctly.
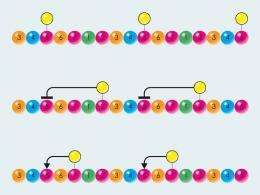
Matthias Geyer and his team at the Max Planck Institute of Molecular Physiology in Dortmund have tracked the functioning of such an enzyme, the transcription factor P-TEFb. P-TEFb equips the reading enzyme RNA polymerase II with phosphate messages. These are reversibly attached to the C-terminal domain (CTD) and play an important role in ensuring that a copy of the base sequence can be produced. The researchers discovered the pattern, according to which P-TEFb attaches the phosphate residues, as well as how existing marks influence the activity of the enzyme.
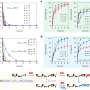
“As we discovered, P-TEFb can not generate certain phosphorylation patterns in the form in which they are described in the literature,” explains Matthias Geyer. The reason for this is that it would appear that some marks influence each other: if a certain position is already marked, another must remain empty – just like an online form in which it is only possible to click on one box. As a result of this discovery, the researchers refuted an accepted textbook principle: “Up to now, it was assumed that, within a repeat unit of seven amino acids, P-TEFb can phosphorylate the serines at positions two and five,” explains the scientist. “In reality, however, the enzyme cannot phosphorylate position two at all, and it can only phosphorylate position five if position two does not yet contain a phosphate residue. The enzyme is also not able to generate a combination of phosphate residues at positions two and five under any circumstances.” Another finding that surprised the researchers was that the enzyme increases its activity by a factor of four if the serine in position seven is already pre-phosphorylated. However, they do not yet know why this is the case.
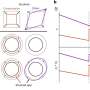
The CTD is an old component of RNA polymerase II in evolutionary terms. In humans, this area consists of 52 sequential repetitions of a motif comprising seven amino acids. Most of these amino acids can be equipped with phosphate attachments, which provide wide-ranging possibilities for the coding and dissemination of information. The researchers assume that messages can even be exchanged in this way between the reading enzyme and the histones – the packaging proteins that act as reels, on which the thread-like genetic substance is wound and stored in the cell nucleus. The histones also carry small chemical attachments and scientists refer to a separate “histone code”. During the reading process, the histones lie in close proximity to the CTD and this facilitates the flow of information.
The molecular messages that P-TEFb introduces within the C-terminal domain are not only important for gene regulation in the healthy organism, but also play an important role in a number of diseases, including certain types of cancer, cardiac muscle diseases and HIV. For example, the HIV virus exploits the enzyme for its purposes to accelerate the reading of the genetic code and induce the cell to start producing virus proteins as quickly as possible. Therefore, the researchers would like to understand the functioning of the enzyme in as much detail as possible so that they can find more out about the disease mechanisms.
The study of the Dortmund-based scientists is the most detailed carried out up to now on the functioning of P-TEFb. The researchers succeeded in analysing the phosphate patterns in detail with the help of genetically modified RNA polymerase II molecules, which they transposed with defined volumes of the enzyme. They were then able to determine precisely which positions within the CTD were provided with phosphate marks, and how the activity rate of the enzyme.
More information: Nadine Czudnochowski, et al., Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition, Nature Communications 3:842 (May 15, 2012), doi: 10.1038/ncomms1846
Journal information: Nature Communications
Provided by Max Planck Society















