This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
First molecular insights into the degradation of the ribosomal 30S subunit
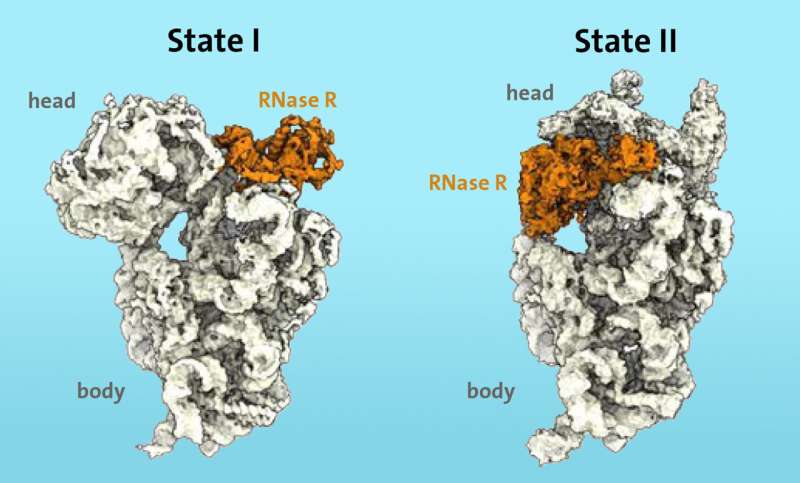
A research team from the Department of Chemistry at the Universität Hamburg has succeeded for the first time in identifying at the molecular level the dynamic mechanism used by the enzyme RNase R to degrade the ribosomal 30S subunit. The results of the study were published in the scientific journal Nature.
Protein synthesis is a vital and energy-intensive process in the cell in which ribosomes play a crucial role. These comparatively large molecules are found in all living organisms and act as the cell's "protein factories."
To do this, ribosomes read the blueprint for a specific protein on a messenger molecule—the messenger RNA (mRNA)—and then convert this information into a new protein. Ribosomes consist of two subunits. The small subunit is responsible for reading and checking the mRNA for errors, while the large subunit is responsible for the polymerization of amino acids to form proteins.
Controlled production and regulated turnover of ribosomes is required for protein synthesis. While the assembly of ribosomes has been increasingly better understood in recent years, there has been no structural insight into the degradation of ribosomes.
This is important because in stress situations such as a lack of food, or at the end of their growth cycle, cells reduce their metabolism in order to survive longer. This state is known as the stationary phase. During this phase, energy-intensive protein synthesis is reduced, and some ribosomes are degraded in order to release the energy invested in them to ensure cell survival.
For their investigations, the researchers studied Bacillus subtilis, a rod-shaped soil bacterium that is found in air, dust and water as well as in the intestines of humans and animals. "In contrast to previous studies, we took cells that were still growing and not in the stationary phase. We wanted to know what processes take place at the transition to the stationary phase," says Dr. Helge Paternoga from the Department of Chemistry at the Universität Hamburg, last author of the study.
The researchers knew from previous work that certain enzymes, such as ribonuclease R (RNase R), are involved in the degradation process of ribosomes in stress situations. Using cryo-electron microscopy, they were able to show for the first time that the enzyme RNase R binds to the small 30S subunit of the ribosome. The "S" stands for "Svedberg units" and refers to the mass of the ribosomal subunit.
The RNase R does not arbitrarily cut the 30S subunit, but rather attaches itself to a free area, which the researchers call the "neck," and then detaches the "head," the upper area of the subunit, in two consecutive stages.
"In the first stage, the enzyme RNase R encounters an obstacle at the 'neck' and destabilizes the neck area, making it more flexible. In the second stage, the 'head' is turned, which removes the obstacle and allows the enzyme to continue the degradation process of the 30S subunit unhindered," explains Paternoga.
"Our in-vitro degradation experiments indicate that the 'head' switch is a significant kinetic barrier for RNase R. Moreover, we were able to show that the enzyme alone is sufficient to accomplish the complete 30S degradation process" says Prof. Dr. Daniel Wilson, head of the research group at the Department of Chemistry at the Universität Hamburg and co-author of the study.
More information: Lyudmila Dimitrova-Paternoga et al, Structural basis of ribosomal 30S subunit degradation by RNase R, Nature (2024). DOI: 10.1038/s41586-024-07027-6
Provided by University of Hamburg