This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Microbial comics: RNA as a common language, presented in extracellular speech-bubbles
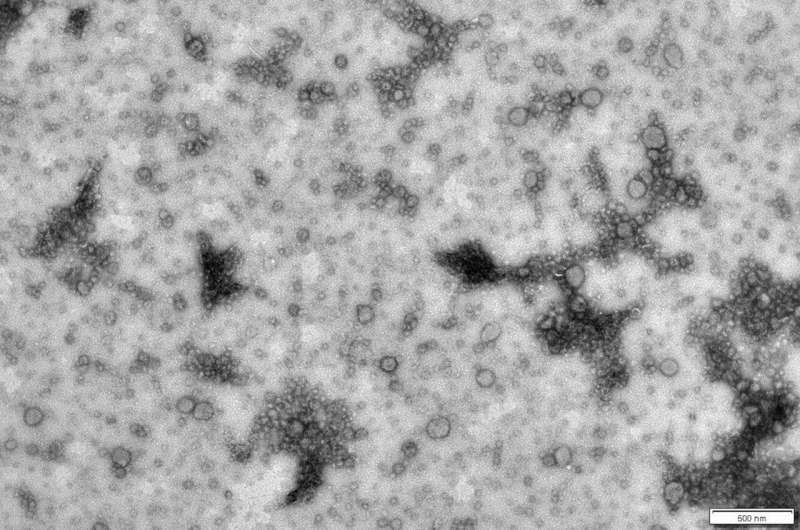
Single-celled organisms, such as bacteria and archaea, have developed many ways to communicate with each other. For example, they might use tiny so-called extracellular vesicles (EVs)—membrane-enveloped packages smaller than 200 nm in diameter (0.0002 mm). The organisms produce them by budding from their membrane into the surrounding space.
These EVs can contain a variety of molecules such as enzymes, nutrients, RNA and even fragments of DNA. Though it is suspected that they play a key role in microbial communities, little is known about their function or how they are produced.
Speech balloons for RNA talk
In a study now published in the Proceedings of the National Academy of Sciences, Susanne Erdmann with her team at the Max Planck Institute for Marine Microbiology and collaborators from other institutions in Germany and Australia investigated EVs from microbes that thrive in extremely salty environments such as the Dead Sea, known as halophilic archaea or haloarchaea.
They found that their EVs traffic RNA—nucleic acids that play a central role in protein synthesis and gene regulation—between cells. "Obviously, EVs can act as an RNA communication system between haloarchaea," Erdmann explains. In particular, the EVs transported specific RNAs with the potential to regulate processes in a receiving cell.
"We think that this represents a communication mechanism to regulate gene expression across a whole microbial population. One could say, RNA is their common language, and the EV is the speech balloon," says Erdmann.
A GTPase known from eukaryotic cells
The team around Erdmann also investigated how the haloarchaea produce these EVs. "We found a small GTPase—a class of enzymes serving as molecular switches or timers in many fundamental cellular processes—that was very similar to a GTPase in more complex cells," reports first-author Joshua Mills, who conducted the study as party of his doctoral thesis.
"That is quite astonishing, as GTPase-dependent vesicle formation was previously thought to be only carried out within eukaryotic cells, between the membrane-bound intracellular compartments. Our finding suggests that components of eukaryotic intracellular vesicle trafficking could have evolved much earlier in evolutionary history than previously assumed."
"Few studies have investigated the role of EVs within the archaeal domain to date," Erdmann adds. "Here we show that EVs in salt-loving archaea can transport an RNA cargo and thus help cells communicate with each other. Also, we reveal exciting new insights into the evolutionary development of this communication strategy.
"Our study provides the basis for further studies into the evolutionary relationships between prokaryotic and eukaryotic vesicle formation and might help solving the puzzle of the evolution of the eukaryotic cell."
More information: Joshua Mills et al, Extracellular vesicle formation in Euryarchaeota is driven by a small GTPase, Proceedings of the National Academy of Sciences (2024). DOI: 10.1073/pnas.2311321121
Journal information: Proceedings of the National Academy of Sciences
Provided by Max Planck Society