This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
Analyses provide new insights into the evolution, domestication and ornamental traits of crape myrtle
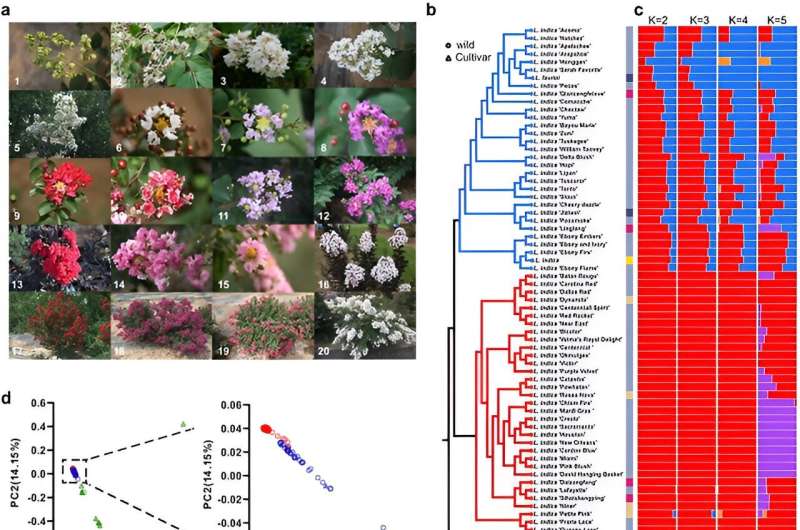
Crape myrtle (Lagerstroemia indica), a widely cherished ornamental plant, boasts a rich history, originating in Southeast Asia to Oceania and flourishing in cultivation centers like China for over 1,600 years. Renowned for its unique blooming during the summer peak, it has evolved through extensive hybridization, and now includes more than 200 species.
Current research has made strides in understanding the determinants of plant architecture, flower, leaf color, and dwarfism traits through transcriptomics and QTL mapping. However, the absence of a reference genome for L. indica severely limits comprehensive genomic studies. This gap hinders the exploration of molecular mechanisms underlying key traits and impedes advancements in plant evolution, domestication, and genomic breeding.
In July 2023, Horticulture Research published a research article titled "Genome assembly and resequencing analyses provide new insights into the evolution, domestication and ornamental traits of crape myrtle ."
In this study, the first high-quality genome assembly of Lagerstroemia indica (L. indica) was performed utilizing PacBio sequencing and Hi-C scaffolding.
The genome annotation identified 33,608 genes and around 42.19% of the genome as repetitive elements. Comparative genomic analysis with 17 species mapped gene families, uncovering 3,572 genes unique to L. indica.
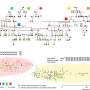
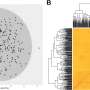
Evolutionary analysis revealed a recent whole-genome triplication in L. indica, and a phylogenetic tree positioned L. indica with L. speciosa and P. granatum in Myrtales. Population structure analysis of 73 accessions using SNPs revealed the evolutionary history and breeding of modern crape myrtle cultivars.
Principal component analysis and pairwise sequential Markovian coalescent showed rich polymorphism among species, with distinct fluctuations in population size. The study of linkage disequilibrium and Tajima's D values highlighted the increased genetic diversity in cultivar accessions.
A high-density genetic linkage map for L. indica was created using an F1 population of L. fauriei and L. indica 'Pocomoke'. This map contained 5,660 SNP markers divided into 24 linkage groups. QTL analysis of plant architecture traits identified 33 intervals affecting phenotypic variation, with a major-effect interval on LG1 significantly influencing internode length.
For flower color, transcriptome sequencing of flowers of different colors identified a large number of DEGs involved in flavonoid pathways essential for petal color determination. Additionally, bulk segregant analysis mapped leaf color loci to chr12 and chr17, and identified five candidate genes, including homologs to MYB35, NCED, and KAS1, related to pigmentation.
In conclusion, this study not only elucidates L. indica's genome and evolutionary history but also provides insights into the genetic basis of its ornamental traits, offering a foundation for future research and molecular breeding in Myrtaceae.
More information: Yang Zhou et al, Genome assembly and resequencing analyses provide new insights into the evolution, domestication and ornamental traits of crape myrtle, Horticulture Research (2023). DOI: 10.1093/hr/uhad146
Journal information: Horticulture Research
Provided by Plant Phenomics