This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Engineered enzymes could generate biomass optimized for conversion into fuel and other useful products
Plant biologists at the U.S. Department of Energy's (DOE) Brookhaven National Laboratory have engineered enzymes to modify grass plants so their biomass can be more efficiently converted into biofuels and other bioproducts. As described in a paper published in Plant Biotechnology Journal, these enzymes modify molecules that make up plant cell walls to provide access to fuel-generating sugars normally locked within complex structures.
"The concept of biomass to biofuel seems simple, but it is technically very difficult to release the sugars," noted Chang-Jun Liu, a senior plant biologist at Brookhaven Lab who led the study.
Plant biomass is full of energy-rich complex sugar molecules generated from photosynthesis. Each plant cell is surrounded by a rigid cell wall made of sugars and a material called lignin that provides structural support. Reducing lignin to gain access to the sugars has been the focus of research aimed at using plants to generate fuels and other products commonly made from petroleum.
For nearly 15 years, Liu has been tackling this problem using engineered enzymes called monolignol 4-O-methyltransferases (MOMTs). These enzymes, which do not exist in nature, are designed to alter the chemical structure of monolignols—the main building blocks of lignin. Changing the structure of the building blocks prevents them from linking together, which reduces the lignin content of plants and makes the sugars more accessible.
In prior work, Liu and his colleagues successfully expressed MOMTs in poplar trees. These enzymes reduced the trees' lignin content and enabled more abundant sugar release from the plants. In the new research, they tested the potential applications of the MOMT enzymes in grass plants, which have an abundant biomass yield.
Grasses can also grow in harsh environments, such as on soils deficient in water or nutrients. Cultivating engineered plants in such environments could potentially produce large amounts of biomass optimized for conversion to fuel and bioproducts—without competing for land needed to produce food crops.
"However, grass plant cell walls, like those from the rice plants that we studied, are even more complicated in terms of structure and composition," explained Nidhi Dwivedi, Brookhaven Lab research associate and lead author on the new paper. In addition to sugar and lignin, grass plant cell walls also contain additional phenolic compounds that "cross-link" the cell wall components, making them even stronger and harder to break down.
"The complexity of grass plant cell walls made us curious as to whether our enzymes would improve sugar recovery," noted Liu. "We wanted to know if MOMTs could modify the grass cell walls in a way that would provide access to the biomass."

Less lignin, more sugars
Liu and Dwivedi chose to focus on two versions of the enzyme for this study—MOMT4 and MOMT9—each of which were designed to modify a different lignin subunit.
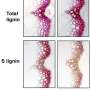
Working with collaborators from Kyoto University in Japan, Liu's team conducted chemical analyses on rice plants engineered to express either MOMT4 or MOMT9. These studies showed there was less lignin in the modified grass plants compared to unaltered plants.
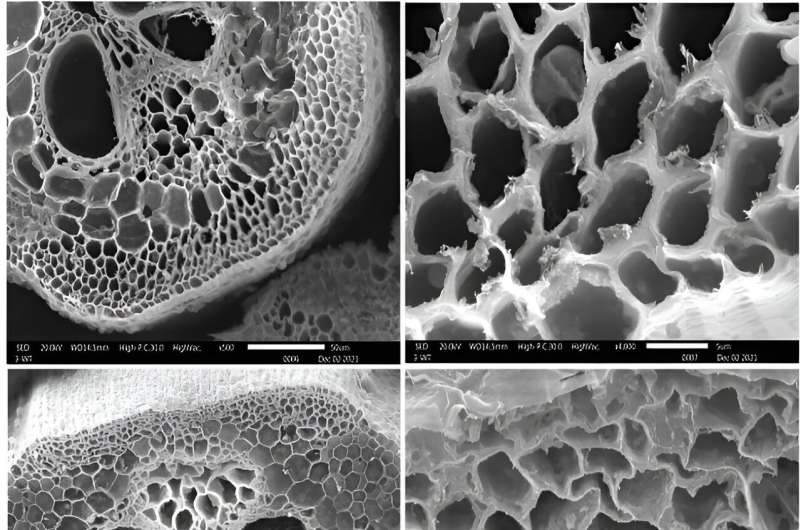
Collaborators from Appalachian State University in North Carolina examined sections of the modified plant stems using scanning electron microscopy and observed changes that were consistent with the chemical analyses.
"Throughout the stem, the cell walls appeared thinner," said Dwivedi. "And in some cells the walls even looked deformed or buckled."
With less lignin in the cell walls, the scientists were able to collect up to 30% more sugar from plants expressing MOMT4 and up to 15% more sugar in plants expressing MOMT9, compared to unaltered plants. Through a process called fermentation, this sugar can be converted into biofuels like ethanol, which is a common additive used to lower the fossil fuel content of gasoline.
Surprisingly promiscuous enzymes
Enzymes—molecules that generally facilitate chemical reactions—commonly target just one type of molecule. MOMT4 and MOMT9 were designed to act upon monolignols. But when Liu and his colleagues ran tests on these enzymes, the results revealed that these engineered enzymes exhibited "promiscuity."
In addition to acting on the monolignols, both MOMTs acted on other cell wall components—the cross-linking phenolics and also a phenolic called tricin, which is a lignin precursor unique to grass plants.
When these enzymes were expressed in rice plants, they made the expected structural changes to the traditional lignin building blocks, and thus reduced the plants' overall lignin content. But by changing the structures of the cross-linking phenolics and tricin, the MOMTs also reduced the incorporation of those compounds into the cell walls further weakening them. The scientists also found an accumulation of modified phenolics in the rest of the plant tissue that was not present in unaltered plants.

"This was quite a difference from what we saw when we expressed the same enzymes in poplar trees," noted Liu. "The broader effects of expressing the enzymes really surprised us. Overall, the changes were positive in terms of optimizing sugar yield from grass cell walls. But there were also some unintended effects."
For example, plants expressing MOMT9 did not grow as tall as the unaltered plants, reducing the quantity of biomass from which sugar could be accessed. The plants also failed to produce seeds, which would be a problem if the scientists want the modified plants to reproduce as a sustainable source of biofuel sugars.
To address these challenges, the scientists plan to explore methods for controlling how lignin gets modified in different parts of the plant. For example, if the scientists can reduce lignin levels everywhere in the plant other than the reproductive organs, they could maximize the ability to extract sugars without affecting the fertility of the plants.
The scientists also want to see if their MOMT enzymes can optimize sugar yields from other grass plant species.
"After seeing the effectiveness of this enzyme technology in rice, we are confident that it can be used to modify other grass energy crops like sorghum and bamboo," Liu said.
"Biofuels are a promising alternative to non-renewable energy sources," Dwivedi added, "This study provides insights into how scientists can optimize the release of sugar that is present in cell walls, thus overcoming some of the waste that occurs with unmodified biomass crops."
More information: Nidhi Dwivedi et al, Simultaneous suppression of lignin, tricin and wall‐bound phenolic biosynthesis via the expression of monolignol 4‐O‐methyltransferases in rice, Plant Biotechnology Journal (2023). DOI: 10.1111/pbi.14186
Journal information: Plant Biotechnology Journal
Provided by Brookhaven National Laboratory