This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Study clarifies how 'junk DNA' influences gene expression
For decades, scientists have known that, despite its name, "junk DNA" in fact plays a critical role: While the coding genes provide blueprints for building proteins, which direct most of the body's functions, some of the noncoding sections of the genome, including regions previously dismissed as "junk," seem to turn up or down the expression of those genes.
But it's been unclear how certain noncoding regions influence gene-expression levels—that is, the number of times a gene is copied into RNA and used to make proteins.
Now, a new study by Polly Fordyce, Ph.D., associate professor of bioengineering and of genetics, and her colleagues has unraveled some of the mystery. Their discovery may help researchers understand complex genetic conditions, including autism, schizophrenia, cancer and Crohn's disease.
"We've known for a while that short tandem repeats, or STRs, aren't junk because their presence or absence correlates with changes in gene expression," Fordyce said. "But we haven't known how they exert these effects."
Authors of the study, published Sept. 22 in Science, believe it's the first to offer a roadmap to understanding how STR changes can impact gene expression.
An evolving view of 'junk DNA'
STRs make up about 5% of the human genome. "Starting in the 1980s, researchers noticed that changes to these repetitive sequences can affect gene expression," said the study's lead author, Connor Horton, who was a technician in Fordyce's lab. "That's the trail of breadcrumbs we've followed."
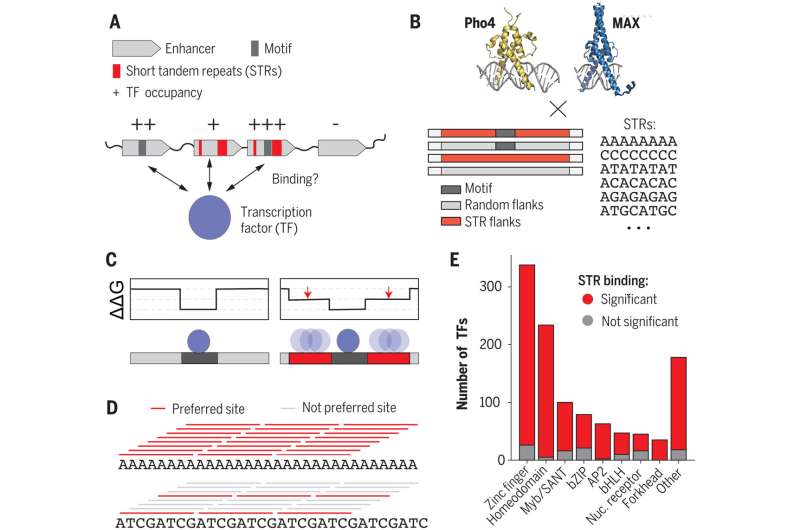
For the study, the researchers looked at how STRs interact with proteins called transcription factors. Transcription factors attach to noncoding DNA, regulating the expression of protein-coding genes.
"Researchers have spent a lot of time characterizing these transcription factors and figuring out which sequences—called motifs—they like to bind to the most," Fordyce said. But current models don't adequately explain where and when transcription factors bind to noncoding DNA to regulate gene expression. Sometimes, no transcription factor is attached to something that looks like a perfect motif. Other times, transcription factors bind to stretches of DNA that aren't motifs.
"To solve the puzzle of why transcription factors go to some places in the genome and not to others, we needed to look beyond the highly preferred motifs," Fordyce said. "In this study, we're showing that the STR sequence around the motif can have a really big effect on transcription factor binding, providing clues as to what these repeated sequences might be doing."
To better understand the role of short tandem repeats in gene expression, the researchers stripped the mechanisms down to their basics: transcription factors and naked DNA. They used specialized assays designed in the Fordyce lab to run thousands of tiny experiments side by side, saving time and money.
The experiments compared how tightly transcription factors attached to thousands of DNA sequences—those with a preferred motif, those without one, and those surrounded by random sequences or by a wide variety of STRs.
"In the experiment we asked, 'How do these changes impact the strength of transcription factor binding?'" Fordyce said. "We saw a surprisingly large effect. Varying the STR sequence around a motif can have up to a 70-fold impact on the binding."
To discover how the DNA and transcription factors interacted, Horton made hundreds of mutated transcription factors. He saw that changes to the transcription factor's DNA binding domain affected whether it recognized the motif and the STRs. The researchers concluded that the transcription factors directly interact with the repetitive genetic code, attaching to it and the motif with the DNA binding domain.
Models to help understand polygenic diseases
The large number —over 6,000—of experiments the team ran made it possible to develop a model of the rules governing transcription-factor binding. Their findings could even help researchers understand and model interactions between other transcription factors and noncoding regions of DNA that regulate gene expression.
"We set out to study short tandem repeats. But the models we developed apply broadly to the entire regulatory landscape," said Horton, now a graduate student at the University of California, Berkeley. "It helps us better understand how transcription factors bind to regulatory DNA, even when short tandem repeats aren't involved."
Models of how noncoding regulatory regions impact transcription factor binding can help researchers understand the role of these sequences in polygenic diseases. "It's been known for some time that short tandem repeats are associated with increased or decreased risk of certain diseases," Horton said.
"We hypothesize that changes in the short tandem repeats between individuals lead to different amounts of transcription factor binding, which leads to changes in gene expression, which might be linked to these diseases."
Through the years, genome-wide association studies have linked changes in STRs to various diseases. "But it wasn't clear what to do with that information," Horton said. "Our models can suggest experiments to understand how those short tandem repeats affect progression or risk of the disease."
More information: Connor A. Horton et al, Short tandem repeats bind transcription factors to tune eukaryotic gene expression, Science (2023). DOI: 10.1126/science.add1250
Journal information: Science
Provided by Stanford University Medical Center