This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A better understanding of gene regulation in embryonic stem cells
During the first hours of an embryo's development, specialized molecules called pioneer transcription factors unravel parts of its DNA to activate the encoded genes. Which gene is activated and when has to follow a set schedule so that genes that are only needed at later stages of development are not activated too early—such as those that trigger the differentiation of specialized cell types.
A previously unknown mechanism that enables this precise control of genes has been discovered by a research team led by developmental biologist Dr. Daria Onichtchouk and theoretical physicist Prof. Dr. Jens Timmer. Both scientists are members of the Cluster of Excellence CIBSS–Center for Integrative Biological Signaling Studies at the University of Freiburg, which promotes interdisciplinary projects like this.
The current study led by the two scientists describes a previously unknown mechanism through which two activating transcription factors block each other's activities. The same factors that trigger the expression of early genes can thereby also prevent the premature expression of later genes. The findings have been published in Nature Communications.
Like the interaction of two keys and a lock
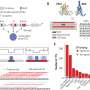
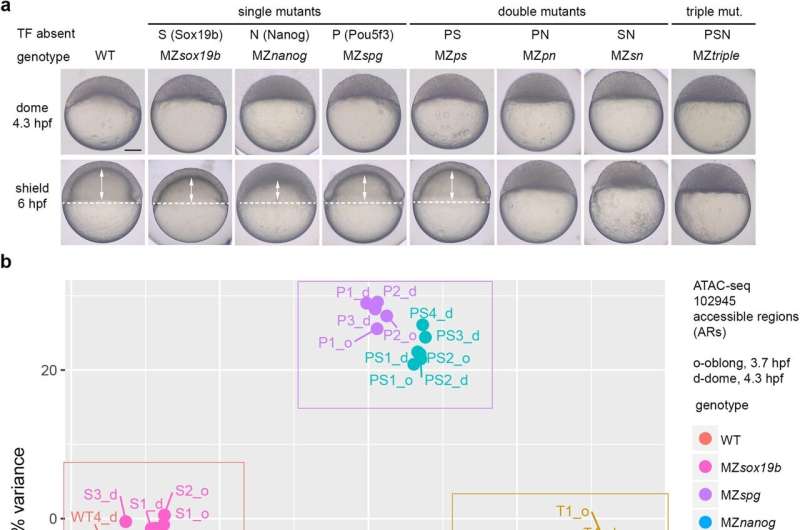
The researchers discovered this in two zebrafish transcription factors, Pou5f3 and Nanog. "We knew that these transcription factors interact, but it was assumed that they could only make DNA more accessible, not less," explains Onichtchouk. The researchers have now been able to show that there are indeed two possible modes of interaction: Pou5f3 and Nanog can either function synergistically to activate a gene, or antagonistically, inhibiting gene expression.
"How they interact depends on the sequence of the DNA region they bind to," says Onichtchouk. "This newly-discovered antagonistic interaction is what inhibits the expression of hundreds of genes during early stages of development."
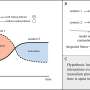
The Freiburg researchers' study shows that Pou5f3 and Nanog synergistically activate those genes where they both act as activators, but have an antagonistic regulatory effect for those genes that can be activated by only one of them. In their publication, the researchers compare this interaction to two keys and a lock: If one of the keys fits into the lock but cannot turn it, it will block the correct key from entering.
"Now we know a lot more about the mechanisms by which transcription factors have context-dependent function," Onichtchouk summarizes their finding. "This adds to the known complexity of gene regulation, but also brings us an important step closer to a complete understanding."
Interdisciplinary approach, simple and general formulas
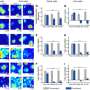
To reach this conclusion, the research groups combined their expertise in developmental biology, genetics and mathematical modeling. Onichtchouk's research group first compared the transcription factors' effect on the expression of several hundreds of genes, using genetic assays. Then Timmer's group used the data to model possible regulation mechanisms, without any further details on the specific factors they were investigating.
Jacques Hermes, who is doing his Ph.D. in Timmer's group and did a large part of the modeling for this study, described the scientific procedure. "The formulas we used were actually very simple and general. That means that the formulas have the same structure for any transcription factor, while the values of the parameters in the formulas are different for every transcription factor combination and every target. Estimating these parameters is a central task in our daily work and in this project we even had a way to immediately validate the results yielded by our model using independent experimental data."
The mammalian counterparts of these molecules, Oct4 and Nanog, are of high medical relevance because they are essential for the induction of induced pluripotent stem cells. Artificial production of such pluripotent stem cells from body cells is a technology with high potential for future use in regenerative medicine which was recognized with the 2012 Nobel Prize.
More information: Aileen Julia Riesle et al, Activator-blocker model of transcriptional regulation by pioneer-like factors, Nature Communications (2023). DOI: 10.1038/s41467-023-41507-z
Journal information: Nature Communications
Provided by University of Freiburg