This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Getting protein factories to run—How deubiquitinating enzymes moonlight as Fubi proteases
The small protein ubiquitin is particularly famous for marking proteins for degradation but it has also been shown to regulate virtually all cellular processes. In parallel to the ubiquitin system various other ubiquitin-like modifiers have evolved, of which Fubi is particularly poorly studied despite its immunomodulatory activity.
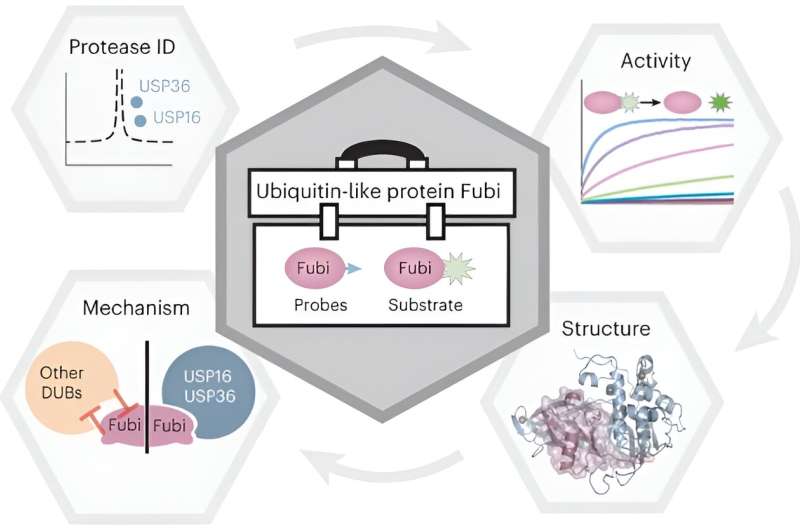
Scientists around Malte Gersch, research group leader at the Chemical Genomics Centre at the Max Planck Institute of Molecular Physiology, have now gained first molecular insights into the machinery facilitating the Fubi-controlled maturation of a key protein of the ribosome, the cell's protein factory. With the help of a newly developed chemical tool kit, the researchers characterized how two deubiquitinating enzymes provide specific Fubi hydrolase activity and thereby moonlight as Fubi proteases in a two-tier manner.
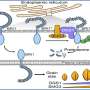
Fubi is produced by cells as a fusion protein with the ribosomal protein S30, and must be separated from S30 by proteases for functioning ribosomes. In immune cells, this by-product of ribosome production is utilized as a secreted signaling molecule, for example to locally reduce the activity of the maternal immune system in the uterus and to thus enable embryos to implant. How Fubi is specifically recognized by proteases and how they distinguish it from ubiquitin was previously unknown.
Rachel O'Dea and Malte Gersch explain their research in detail:
What is the discovery that you made and why is it exciting?
Our team revealed how two deubiquitinating enzymes can also act as proteases of the ubiquitin-like protein Fubi and gained molecular insights into how this is possible in specific manner. This is noteworthy because, despite similarity between Ubiquitin and Ubiquitin-like proteins, the enzymes regulating them in humans are usually not the same.
We show that this dual activity is specific to the two enzymes USP16 and USP36 and our crystallography studies mechanistically explain how this rare cross-reactivity is achieved. Surprisingly, unlike what is observed in cross-reactive enzymes from bacteria or viruses, we did not find any additional structural elements that facilitate the additional Fubi activity of these well characterized Ubiquitin proteases.
Instead, Fubi recognition is mediated through a small cryptic motif on a complementary binding surface.
What is so special about your developed tool kit?
Possibly due to its challenging amino acid composition Fubi protein tools had not yet been added to the repertoire of tools for studying Ubiquitin and Ubiquitin-like proteins.
Our work demonstrates facile approaches for making Fubi tools which can be readily adapted by other scientists in the field. The probes and Fubi fluorescent substrate described here provide a means for assessing Fubi eraser activity in both cellular and in-vitro settings.
Why is your research important for the scientific community?
Our work provides new molecular insights into how enzymes can have activities spanning multiple modification systems. Explaining how USP16 and USP36 play a role in ribosomal protein maturation expands our understanding of mechanisms regulating this critical cellular process.
Fubi has primarily been studied by scientists from the immunology field, and more recently of the ribosome field, and our work approaching the topic with the Ubiquitin background complements these other works. Together all data converges into a two-tier model for Fubi processing.
Why is your research important for society?
Owed to their rapid and reversible nature posttranslational modifications such as Ubiquitin and Ubiquitin-like proteins are critical regulators of virtually all cellular processes.
Fubi has been linked to immunomodulatory functions and has been shown to modify proteins during immune stimulation responses. Understanding the exact role of Fubi in this process will expand our understanding of the how cells respond to immune signaling.
What are the next steps you will take?
Our insights into Fubi recognition allow for tuneable Fubi protease activity in cells and are thus paving the way for better understanding the cellular role of this enigmatic protein as a post-translational modification.
In addition, we are using the probes to facilitate investigation into the molecular mechanism by which other proteins interact with Fubi. But first we will celebrate.
The findings are published in the journal Nature Chemical Biology.
More information: Rachel O'Dea et al, Molecular basis for ubiquitin/Fubi cross-reactivity in USP16 and USP36, Nature Chemical Biology (2023). DOI: 10.1038/s41589-023-01388-1
Journal information: Nature Chemical Biology
Provided by Max Planck Society