This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers uncover how photosynthetic organisms regulate and synthesize ATP
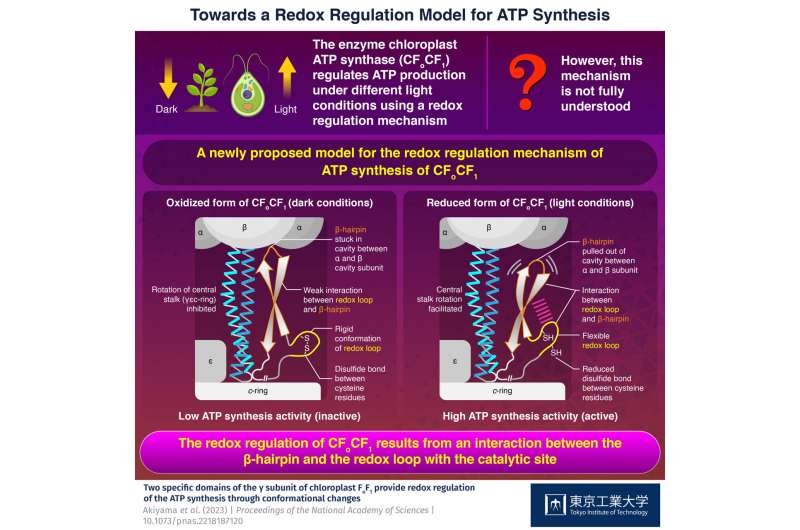
ATP, the compound essential for the functioning of photosynthetic organisms such as plants, algae, and cyanobacteria, is produced by an enzyme called "chloroplast ATP synthase" (CFoCF1). To control ATP production under varying light conditions, the enzyme uses a redox regulatory mechanism that modifies the ATP synthesis activity in response to changes in the redox state of cysteine (Cys) residues, which exist as dithiols under reducing (light) conditions, but forms a disulfide bond under oxidizing (dark) conditions. However, this mechanism has not yet been fully understood.
Now, in a study published in the Proceedings of the National Academy of Sciences, a team of researchers from Japan led by Prof. Toru Hisabori from Tokyo Institute of Technology (Tokyo Tech) has uncovered the role of the amino acid sequences present in CFoCF1, revealing how the enzyme regulates ATP production in photosynthetic organisms.
To understand how the conformation of the amino acids present in CFoCF1 contributes to the redox regulation mechanism, the researchers used the unicellular green alga, Chlamydomonas reinhardtii, to produce the enzyme. "By leveraging the powerful genetics of Chlamydomonas reinhardtii as a model organism for photosynthesis, we conducted a comprehensive biochemical analysis of the CFoCF1 molecule," explains Prof. Hisabori.
With the alga as the host organism, the team introduced plasmids (extrachromosomal DNA molecule that can replicate independently) that encoded the F1 component of the CFoCF1 protein, namely the part of the enzyme containing catalytic sites for ATP synthesis. They additionally introduced mutated versions of the gene to change the amino acid sequences of the protein, specifically targeting the DDE motif (a cluster of negatively charged amino acids), the redox loop, and the β-hairpin domain.
They then purified CFoCF1, generating five different variations of it that included a wild-type strain with no changes to the amino acid sequence and four mutant strains: one with the DDE motif replaced with neutral amino acids, Asn-Asn-Gln, one without the β-hairpin domain, one without the redox loop, and one lacking both the redox loop and the β-hairpin domain.
Upon testing the ATP synthesis activity of these mutants under reducing (mimicking the light conditions) and oxidizing (mimicking the dark conditions) conditions, the researchers found that the wild-type enzyme and the mutant enzyme with changes to the DDE motif functioned normally (showed high activity when reduced and low activity when oxidized). However, the enzyme complexes without the redox loop or the β-hairpin domain did not show the redox-response, indicating that both regions were involved in the redox regulation mechanism.
The researchers suggested that under dark conditions, the disulfide bond between the Cys residues makes the redox loop rigid and weakens the interaction between the redox loop and the β-hairpin. This causes the β-hairpin to remain stuck within a cavity in the protein. However, when the disulfide bond is reduced in presence of light, the redox loop regains its flexibility and pulls the β-hairpin out of the cavity, enabling it to partake in the ATP synthesis activity.
"The redox regulation of ATP synthesis is accomplished by a cooperative interaction between two γ subunit domains of CFoCF1 unique to photosynthetic organisms," says Prof. Hisabori. "We propose that it results from the interaction of the β-hairpin and the redox loop with the catalytic site."
The results are an important step towards understanding the photosynthesis process better, with the potential for significant implications in the fields of agriculture and bioenergy.
More information: Kentaro Akiyama et al, Two specific domains of the γ subunit of chloroplast F o F 1 provide redox regulation of the ATP synthesis through conformational changes, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2218187120
Journal information: Proceedings of the National Academy of Sciences
Provided by Tokyo Institute of Technology