This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
When editing bacteria with CRISPR, less is more
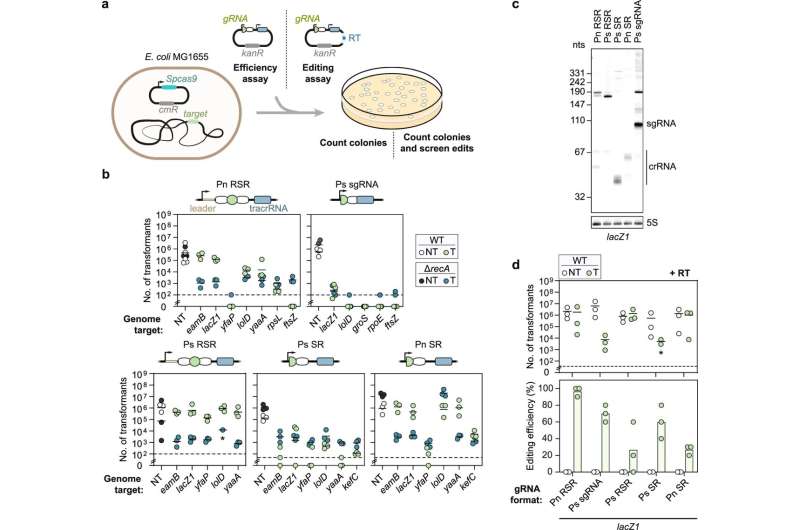
Systematically attenuating DNA targeting activity can achieve CRISPR-driven editing in bacteria, greatly boosting colony counts and even increasing the frequency of precise genome editing. This was shown in a study of the Helmholtz Institute Würzburg (HIRI) in collaboration with the Helmholtz Centre for Infection Research (HZI) in Braunschweig. The findings were published today in the journal Nature Communications.
The ability to genetically manipulate bacteria has been key to exploring the microbial world. Genome editing is crucial for the development of new antibiotics and harnessing bacteria as miniature factories for the sustainable production of chemicals, materials, and therapeutics. Tools based on CRISPR "gene scissors" have proven instrumental in this regard, making it fast, easy, and reliable to create edits in different bacteria.
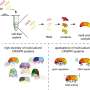
The general technology requires a CRISPR ribonucleic acid (crRNA) serving as a "guide RNA". It helps detect certain regions of a genome for targeted DNA cleavage. Proteins involved in homologous recombination—a natural process of exchanging genetic material between chromosomes—weave in designed "repair template" DNA afterwards to create an edited sequence of the strand.
Breaking the stumbling block
In the current study, researchers from the Würzburg Helmholtz Institute for RNA-based Infection Research (HIRI) in collaboration with the Helmholtz Centre for Infection Research (HZI) in Braunschweig address a central challenge to genome editing in bacteria.
"CRISPR-based genome editing has become a common molecular technology, but there is a notable stumbling block," says HIRI department head Chase Beisel, who led the study. "During exponential growth, bacteria initiate genome replication multiple times in one cell cycle to keep pace with cell division. By efficiently cutting DNA, CRISPR leads to the untimely death of the cell. Consequently, editing requires efficient recombination and high transformation efficiencies, which are unavailable in most bacterial strains, including those relevant to human disease and industrial biotechnology," Beisel explains.
A seemingly paradoxical concept
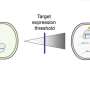
Daphne Collias, postdoc in Beisel's lab at the Helmholtz Institute Würzburg, is the first author of the study. Describing the background to the findings she says, "We found that attenuating the cutting activity of CRISPR would allow the cell to repair the cut DNA using the provided template for homologous recombination. As a result, we could drive homologous recombination and get many more surviving cells."
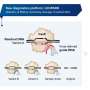
The researchers also developed a suite of approaches that could pull back activity, including using different formats for the guide RNA that directs cutting by the Cas9 protein, using versions of Cas9 that cut less efficiently, reducing guide RNA expression, introducing interfering structures onto the guide RNA, and mutating the sequence in the guide RNA used to find its DNA target.
"We called the modified guide RNAs 'attenuated guide RNAs' or atgRNAs, representing a flexible means of achieving CRISPR-driven editing," Collias reports. "Not every approach could drive editing, although we usually could find at least one for each editing setup."
Future impact
As a proof-of-principle, the Beisel lab teamed up with HZI department head Till Strowig and his lab to enhance editing in different strains of Klebsiella bacteria. Using a multidrug resistant strain, they leveraged editing to reverse resistance against the antibiotic ampicillin.
The new editing approach can advance basic research on bacteria involved in human health. Edited bacteria could also be used as therapeutic probiotics or as production hosts for therapeutics in the future.
More information: Daphne Collias et al, Systematically attenuating DNA targeting enables CRISPR-driven editing in bacteria, Nature Communications (2023). DOI: 10.1038/s41467-023-36283-9
Journal information: Nature Communications
Provided by Helmholtz Association of German Research Centres