This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Antibiotic consumption is currently not the main driver of aminoglycoside resistance spread, study suggests
The spread of antibiotic resistance, whereby infectious bacteria are able to defeat the drugs intended to kill them, may not be primarily driven by antibiotic consumption, according to a study published today in eLife.
Rather, the study suggests that the prevalence of antibiotic resistance across Europe between 1997 and 2018 is mostly explained by exchanges between ecosystems, and human exchanges such as merchandise imports or travel.
The results support the idea that interventional strategies based on reducing antibiotic use should be complemented by a stronger control of exchanges, especially between ecosystems.
Antibiotic resistance represents one of the largest threats to global public health, food security and global development faced today. Due to the spread of antibiotic resistance, a growing number of infections, such as pneumonia and tuberculosis, are becoming harder to treat, leading to longer hospital stays, greater costs and increased mortality.
"Many public health agencies have recommended reducing antibiotic use in response to the challenges caused by resistance," explains co-author Léa Pradier, a former Ph.D. student at University of Montpellier, France. Pradier conducted the study alongside Stéphanie Bedhomme, a researcher at CNRS.
"However, there are cases where developed countries have reduced their antibiotic consumption and not halted the spread of antibiotic resistance genes across bacterial populations, implying other factors are at play," continues Pradier.
To explain this, Pradier and Bedhomme set out to describe the genetic, geographical and ecological distribution of resistances to a class of antibiotics called aminoglycosides, and from this information, quantify the relative contribution of different factors driving the spread of antibiotic resistance.
Aminoglycosides have limited clinical use in humans, but are often a last resort for treating multi-resistant infections. They are also commonly used in the treatment of farmyard animals, meaning that resistance to them poses a significant threat to global food security.
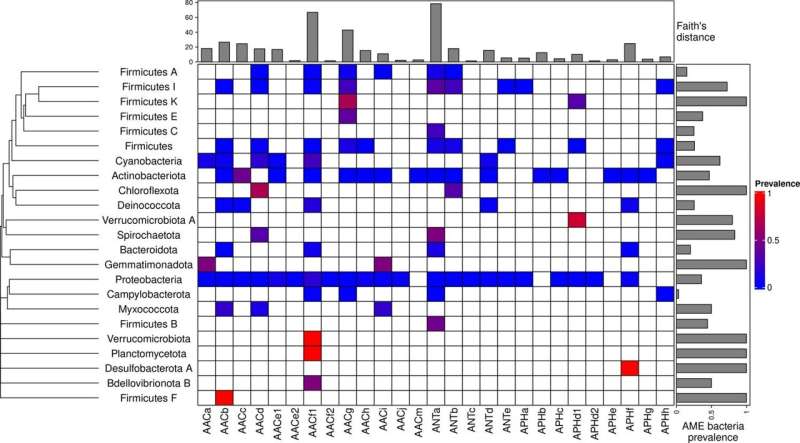
They utilized a computational approach to screen the genetic information of over 160,000 bacteria genomes, looking for genes encoding aminoglycoside-modifying enzymes (AMEs)—the most common mechanism of aminoglycoside resistance. They detected AME genes in around a quarter of genomes screened, and in samples from all continents (excluding Antarctica) and all biomes investigated. The majority of AME-gene-carrying bacteria were found in clinical samples (55.3%), human samples (22.1%) and farm samples (12.3%).
Pradier and Bedhommme then focused on the distribution of AME genes across Europe, from 1997–2018, when the most detailed data was available. During this period, aminoglycoside usage remained relatively constant, but was highly variable between countries. Comparing the prevalence of AME genes between countries with different aminoglycoside usage over time, the team determined that aminoglycoside consumption was only a minor explanatory factor, with few positive or directional effects on AME gene prevalence.
Instead, the dataset implies that human exchanges through trade and migration, and exchanges between biomes, explain most of the spread and maintenance of antibiotic resistance when modeled over time, space and ecology. AME genes can be carried over continents by plant and animal products, and international trade and travelers, and may then spread to local strains of bacteria through a process called horizontal gene transfer—the movement of genetic information between organisms.
The pool of AME genes sampled from plants, wild animals and soil had the strongest overlap with other communities, suggesting these biomes are major hubs for AME gene propagation, either by horizontal resistance gene transfer or by resistant bacteria movement.
The findings suggest that the largest cause of AME gene spread is through the movement of antibiotic-resistant bacteria between ecosystems and biomes. This spread is aided by mobile genetic elements, which increase the likelihood for a genome to carry several copies of the same AME gene. This increases the expression of transferred AME genes and allows bacteria to evolve new antibiotic resistance functions through the duplicated sequences.
These findings are preliminary, as limited by the use of publicly available data, rather than deploying a dedicated sampling method. In addition, the genetic data sourced from multiple different research projects caused a sampling bias towards industrialized countries and biomes with clinical interest, leading to some locations and biomes being over-represented.
"Our study provides a broad overview of the spatial, temporal and ecological distributions of AME genes, and establishes that the recent variations of AME bacteria in Europe are first explained by ecology, then human exchanges and lastly by antibiotic consumption," concludes Bedhomme. "Although the conclusions of this study should not be extended to antibiotic genes other than AMEs, the methods used could easily be applied to further studies on other antibiotic resistance gene families."
More information: Léa Pradier et al, Ecology, more than antibiotics consumption, is the major predictor for the global distribution of aminoglycoside-modifying enzymes, eLife (2023). DOI: 10.7554/eLife.77015
Journal information: eLife
Provided by eLife