This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
An open-source stopwatch to time interactions between molecules inside living cells
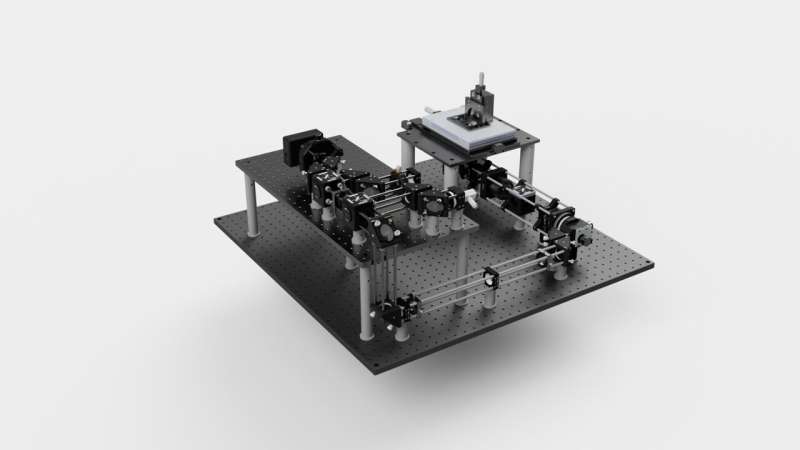

A stopwatch to investigate what happens inside living cells at a thousandth of a millisecond scale: this is the open-source platform BrightEyes-TTM developed by the research team led by Giuseppe Vicidomini at Istituto Italiano di Tecnologia (IIT-Italian Institute of Technology).
BrightEyes-TTM is the first instrumentation to study how molecules inside living cells behave over time, and it can be applied to study variations that occur when a healthy cell becomes diseased. The technology has been published in Nature Communications and in the future, it can be replicated, following the "open science" approach, from other researchers or any maker passionate about technology.
Visualizing and studying the movement of molecules inside cells is challenging because of their microscopic size. Using the optical microscope is not enough if a technology able to distinguish a specific molecule from the thousands of different molecules inside cells is not available.
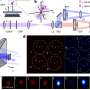
The IIT team exploits the principle of fluorescence, which is the property of specific molecules to become bright when hit by light. By using a special sensor able to record the light at the photon level, so that the light signals emitted by the molecules are recorded providing information about the position and number of molecules inside cells.
The sensor, called a "SPAD array detector," allows obtaining images with more details and higher contrast. It records the turning on and off of fluorescent molecules when hit by light. Researchers focused then on the time between absorbing the light and emitting the fluorescence, the so-called lifetime, which can provide additional information, such as details about the structure and the microenvironment of molecules. Exploiting all the properties of light by combining images with lifetime provides a comprehensive view of the function of molecules inside cells.
The lifetime is represented by short times, less than a thousandth of a millisecond. The new platform BrightEyes-TTM is able to log the instant a light particle is emitted and captured by the sensor, acting as a stopwatch for molecular interactions inside cells.
The BrightEyes-TTM development involved a multidisciplinary team of engineers, physicians, and biologists, under the support of the European Research Council, Fondazione San Paolo, and the European Union's Horizon 2020 program.

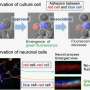
Researchers from IIT are already applying the platform BrightEyes-TTM in the context of the "RNA initiative," an IIT-branded initiative focused on RNA that combines labs and researchers with different expertise. Indeed, investigation of the lifetime of proteins and RNA involved in neurodegenerative diseases will elucidate how these two molecular classes interact with each other before and after the pathological processes behind neuronal death.
Being an open-source system, BrightEyes-TTM is a low-cost method to investigate dynamic processes of molecules. It is now accessible to the international scientific community interested in using or adapting it to different sensors. IIT researchers are convinced that sharing the platform with the scientific community will also pave the way to non-biological applications, such as quantum microscopy.
More information: Alessandro Rossetta et al, The BrightEyes-TTM as an open-source time-tagging module for democratising single-photon microscopy, Nature Communications (2022). DOI: 10.1038/s41467-022-35064-0
Journal information: Nature Communications
Provided by Italian Institute of Technology