This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New method precisely locates gene activity and proteins across tissues
A new method can illuminate the identities and activities of cells throughout an organ or a tumor at unprecedented resolution, according to a study co-led by researchers at Weill Cornell Medicine, NewYork-Presbyterian and the New York Genome Center.
The method, described Jan. 2 in a paper in Nature Biotechnology, records gene activity patterns and the presence of key proteins in cells across tissue samples, while retaining information about the cells' precise locations. This enables the creation of complex, data-rich "maps" of organs, including diseased organs and tumors, which could be widely useful in basic and clinical research.
"This technology is exciting because it allows us to map the spatial organization of tissues, including cell types, cell activities and cell-to-cell interactions, as never before," said study co-senior author Dr. Dan Landau, an associate professor of medicine in the Division of Hematology and Medical Oncology and a member of the Sandra and Edward Meyer Cancer Center at Weill Cornell Medicine and a core faculty member at the New York Genome Center.
The other co-senior author was Dr. Marlon Stoeckius of 10x Genomics, a California-based biotechnology company that makes laboratory equipment for the profiling of cells within tissue samples. The three co-first authors were Dr. Nir Ben-Chetrit, Xiang Niu, and Ariel Swett, respectively a postdoctoral researcher, graduate student, and research technician in the Landau laboratory during the study.
The new method is part of a broad effort by scientists and engineers to develop better ways of "seeing" at micro scale how organs and tissues work. Researchers in recent years have made big advances particularly in techniques for profiling gene activity and other layers of information in individual cells or small groups of cells. However, these techniques typically require the dissolution of tissues and the separation of cells from their neighbors, so that information about profiled cells' original locations within the tissues is lost. The new method captures that spatial information as well, and at high resolution.
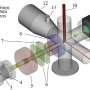
The method, called Spatial PrOtein and Transcriptome Sequencing (SPOTS), is based in part on existing 10x Genomics technology. It uses glass slides that are suitable for imaging tissue samples with ordinary microscope-based pathology methods, but are also coated with thousands of special probe molecules. Each of the probe molecules contains a molecular "barcode" denoting its two-dimensional position on the slide. When a thinly sliced tissue sample is placed on the slide and its cells are made permeable, the probe molecules on the slide grab adjacent cells' messenger RNAs (mRNAs), which are essentially the transcripts of active genes. The method includes the use of designer antibodies that bind to proteins of interest in the tissue—and also bind to the special probe molecules. With swift, automated techniques, researchers can identify the captured mRNAs and selected proteins, and map them precisely to their original locations across the tissue sample. The resulting maps can be considered alone, or compared to standard pathology imaging of the sample.
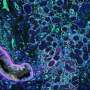
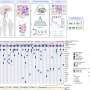
The team demonstrated SPOTS on tissue from a normal mouse spleen, revealing the complex functional architecture of this organ including clusters of different cell types, their functional states, and how those states varied with the cells' locations.
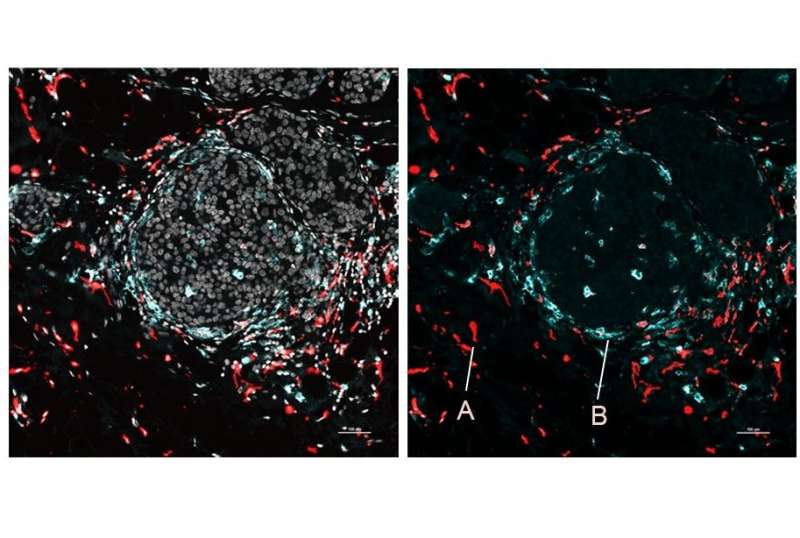
Highlighting SPOTS' potential in cancer research, the investigators also used it to map the cellular organization of a mouse breast tumor. The resulting map depicted immune cells called macrophages in two distinct states as denoted by protein markers—one state active and tumor-fighting, the other immune-suppressive and forming a barrier to protect the tumor.
"We could see that these two macrophage subsets are found in different areas of the tumor and interact with different cells—and that difference in microenvironment is likely driving their distinct activity states," said Dr. Landau, who is also an oncologist at NewYork-Presbyterian/Weill Cornell Medical Center.
"Such details of the tumor immune environment—details that often can't be resolved due to immune cells' sparseness within tumors—might help explain why some patients respond to immune-boosting therapy and some don't, and thus could inform the design of future immunotherapies," he added.
"This initial version of SPOTS has a spatial resolution such that each "pixel" of the resulting dataset sums gene activity information for at least several cells. However, the researchers hope soon to narrow this resolution to single cells, while adding other layers of key cellular information," Dr. Landau said.
More information: Marlon Stoeckius, Integration of whole transcriptome spatial profiling with protein markers, Nature Biotechnology (2023). DOI: 10.1038/s41587-022-01536-3. www.nature.com/articles/s41587-022-01536-3
Journal information: Nature Biotechnology
Provided by Weill Cornell Medical College