This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Boosting efficiency of genome editing procedures to modify initially inaccessible DNA sequences
In the course of optimizing key procedures of genome editing, researchers from the department of Developmental Biology / Physiology at the Center for Organismal Studies of Heidelberg University have succeeded in substantially improving the efficiency of molecular genetic methods such as CRISPR/Cas9 and related systems, and in broadening their areas of application.
Together with colleagues from other disciplines, the life scientists fine-tuned these tools to enable, inter alia, effective genetic screening for modeling specific gene mutations. In addition, initially inaccessible DNA sequences can now be modified. According to Prof. Dr. Joachim Wittbrodt, this opens up extensive new areas of work in basic research and, potentially, therapeutic application.
Genome editing means the deliberate altering of DNA with molecular genetic methods. It is used to breed plants and animals, but also in basic medical and biological research. The most common procedures include the "gene scissors" CRISPR/Cas9 and its variants known as base editors. In both cases, enzymes have to be transported into the nucleus of the target cell. Upon arrival, the CRISPR/Cas9 system cuts the DNA at specific sites, which causes a double strand break. New DNA segments can then be inserted at that site.
Base editors use a similar molecular mechanism but they do not cut the DNA double strand. Instead, an enzyme coupled with the Cas9 protein performs a targeted exchange of nucleotides—the basic building blocks of the genome. In three successive studies, Prof. Wittbrodt's team succeeded in considerably enhancing the efficiency and applicability of these methods.
A challenge when using CRISPR/Cas9 consists in the efficient delivery of the required Cas9 enzymes to the nucleus. "The cell has an elaborate 'bouncer' mechanism. It distinguishes between proteins that are allowed to translocate into the nucleus and those that are supposed to stay in the cytoplasm," explains Dr. Tinatini Tavhelidse-Suck from Prof. Wittbrodt's team. Access is enabled here by a tag made up of a few amino acids that functions like an "admission ticket."
The scientists have now come up with a kind of generally valid "VIP admission ticket" which lets enzymes equipped with it into the nucleus very quickly. They have named it "high efficiency-tag," "hei-tag" for short. "Other proteins that have to penetrate the cell nucleus are also more successful with 'hei-tag,'" says Dr. Thomas Thumberger, who is also a researcher at the Center for Organismal Studies (COS).
In cooperation with pharmacologists from Heidelberg University, the team could show that Cas9 in connection with the "hei-tag" ticket can enable highly efficient, targeted genome alterations not only in the model organism medaka, the Japanese ricefish (Oryzias latipes), but also in mammalian cell cultures and mouse embryos.
In a further study, the Heidelberg scientists showed that base editors operate highly efficiently in the living organism and are even suited to genetic screening. In an experiment with Japanese rice fish, they were able to show that these locally limited, targeted modifications in individual buildings blocks of the DNA achieve an outcome that is otherwise only obtained by the comparatively laborious breeding of organisms with altered genes.
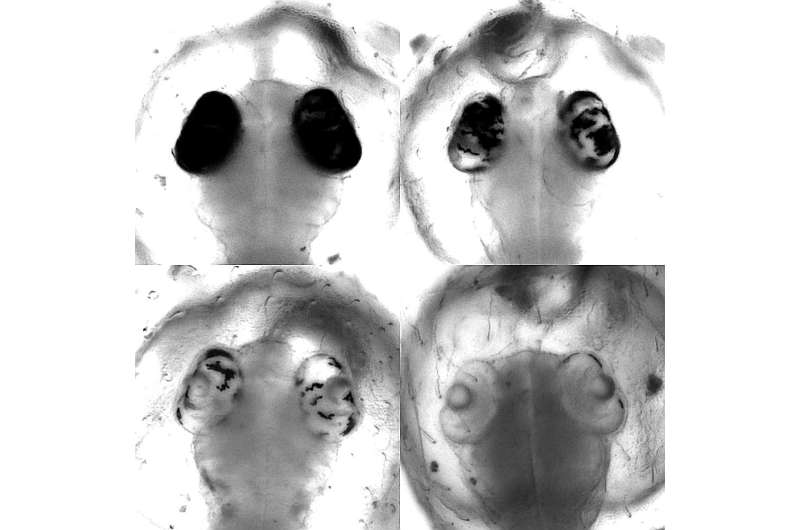
The research team at COS, in cooperation with Dr. Dr. Jakob Gierten, a pediatric cardiologist at Heidelberg University Hospital, focused on certain genetic mutations. These mutations were suspected of triggering congenital heart defects in humans. Through modifying individual building blocks of the DNA of the relevant genes in the model organism, the scientists were able to imitate and study fish embryos with the described heart defects.
The targeted intervention led to visible changes in the heart already during early stages of fish embryonic development, say Bettina Welz and Dr. Alex Cornean, two of the first authors of the study from Prof. Wittbrodt's team. That enabled the researchers to confirm the original suspicion and establish a causal connection between genetic alteration and clinical symptoms.
The precise intervention in the genome of the fish embryos was made possible through especially developed software ACEofBASEs, which is available online. It allows for identifying genetic locations that very efficiently lead to desired changes in the target genes and the resultant proteins. The scientists say that the Japanese ricefish is an excellent genetic model organism for modeling mutations like those identified from the respective patients. "Our method enables an efficient screening analysis and could therefore offer a starting point for developing individualized medical treatment," according to Jakob Gierten.
A third study, again from the Wittbrodt group, deals with the limitations of base editors. For such an editor to bind the DNA of a target cell, there has to be a certain sequence motif. It is called Protospacer Adjacent Motif, PAM for short. "If this motif is lacking near the DNA building block to be changed, it is impossible to exchange nucleotides," explains Dr. Thumberger. A team under his direction has now found a way to get around this limitation.
Two base editors in a single cell are used in succession. In an initial step, a new DNA binding motif for a further base editor is generated, upon which this second editor, which is applied simultaneously, can edit a site that was inaccessible before. This staggered use turned out to be highly efficient, explains Kaisa Pakari, the first author of the study. With this trick, the Heidelberg scientists were able to increase the number of possible application sites of established base editors by 65%. Now DNA sequences that were initially inaccessible can also be modified.
"Optimizing the existing tools for genome editing and their fine-tuned application results in enormously varied possibilities for basic research and, potentially, novel therapeutic approaches," Joachim Wittbrodt underlines.
The research findings were published in the journals eLife and Development.
More information: Thomas Thumberger et al, Boosting targeted genome editing using the hei-tag, eLife (2022). DOI: 10.7554/eLife.70558
Alex Cornean et al, Precise in vivo functional analysis of DNA variants with base editing using ACEofBASEs target prediction, eLife (2022). DOI: 10.7554/eLife.72124
Kaisa Pakari et al, De novo PAM generation to reach initially inaccessible target sites for base editing, Development (2023). DOI: 10.1242/dev.201115
Journal information: Development , eLife
Provided by Heidelberg University