Decoding the cell signals between young proteins and their 'chaperones'
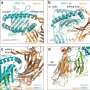
Of the 25,000 different proteins in the human body, insulin, antibodies, and collagen are among the few that perform their biological jobs by literally folding into 3D shapes.
But folding doesn't come easy for newborn proteins. Some get stuck together in the crowded compartment of our cells—the so-called endoplasmic reticulum, or ER—resulting in the production of toxic materials that cause diseases like type 2 diabetes.
To aid their growth, the young proteins enlist the protection of a chaperone called BiP (binding immunoglobulin protein), but how our cells make this match has remained unclear.
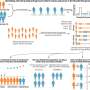
Scientists at the Yale Nanobiology Institute have now decoded the protein signal sequences that determine the movement and timing of the protein-chaperone match—effectively revealing the blueprint for how our proteins reach maturity.
The scholars found that weaker, less hydrophobic signals were associated with a pause in the protein translocation process, triggering the help of the chaperone for entry into the ER compartment and successful protein folding. The stronger, more hydrophobic the signal, the less the need for a chaperone match.
The findings were published today in the Journal of Cell Biology.
Led by Malaiyalam Mariappan, associate professor in the Department of Cell Biology, Yale School of Medicine, the research has the potential to guide the design of customized signal sequences to produce therapeutic recombinant proteins, including insulin, growth factors, and monoclonal antibodies that can be used to treat COVID-19 infection.
More information: Sha Sun et al, Signal sequences encode information for protein folding in the endoplasmic reticulum, Journal of Cell Biology (2022). DOI: 10.1083/jcb.202203070
Journal information: Journal of Cell Biology
Provided by Yale University