Molecular chaperone interactions visualized through X-ray structure analysis
As task forces of the adaptive immune system, T lymphocytes are responsible for attacking and killing infected or cancerous cells. Such cells, like almost all cells in the human body, present on their surface fragments of all the proteins they produce inside. If these include peptides that a T lymphocyte recognizes as foreign, the lymphocyte is activated and kills the cell in question.
It is therefore important for a robust T-cell response that suitable protein fragments are presented to the T lymphocyte. The research team led by Simon Trowitzsch and Robert Tampé from the Institute of Biochemistry at Goethe University Frankfurt has now shed light on how the cell selects these protein fragments or peptides.
Peptide presentation takes place on so-called major histocompatibility complex class I molecules (MHC I). MHC I molecules are a group of very diverse surface proteins that can bind myriads of different peptides. They are anchored in the cell membrane and form a peptide-binding pocket with their outward-facing part.
Like all surface proteins, MHC I molecules take the so-called secretory pathway: they are synthesized into the cell's cavity system (endoplasmic reticulum (ER) and Golgi apparatus) and folded there. Small vesicles then bud off from the cavity system, migrate to the cell membrane and fuse with it.
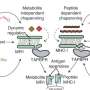
The maturation process of the MHC I molecules is very strictly controlled: in the ER, proteins known as "chaperones" help them fold. The chaperone tapasin is essential for peptide loading in this process.
"When an MHC I molecule has bound a peptide, tapasin checks how tight the binding is," says Trowitzsch, explaining the chaperone's task. "If the bond is unstable, the peptide is removed and replaced by a tightly binding one." However, it has not yet been possible to clarify how exactly tapasin performs this task—especially because the loading process is extremely fast.
The biochemists and structural biologists from Goethe University Frankfurt have now succeeded for the first time in visualizing the short-lived interaction between chaperone and MHC I molecule by means of X-ray structure analysis.
To do this, they produced variants of the two interaction partners that were no longer embedded in the membrane, purified them and brought them together. A trick helped to capture the loading complex in action for crystallization: first, the research team loaded the MHC I molecule with a high-affinity peptide so that a stable complex was created.
A light signal triggered cleavage of the peptide, which greatly reduced its ability to bind the MHC I molecule. Immediately, tapasin entered the scene and remained bound to the MHC I molecule that lacks its peptide. "The photo-induced cleavage of the peptide was pivotal to the success of our experiment," says Tampé. "With the help of this optochemical biology, we can now systematically reproduce complex cellular processes one by one."
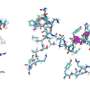
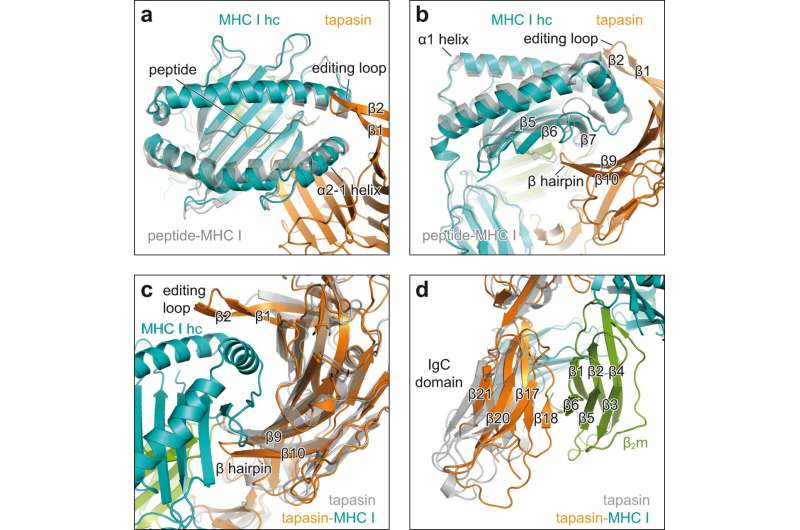
X-ray structure analysis of the crystals revealed how tapasin widens the peptide-binding pocket of the MHC I molecule, thereby testing the strength of the peptide bond. For this purpose, the interaction partners form a large contact area; for stabilization, a loop of tapasin sits on top of the widened binding pocket.
"This is the first time we have shown the process of loading at high resolution," Tampé says. The images also reveal how a single chaperone can interact with the enormous diversity of MHC I molecules, says the biochemist. "Tapasin binds precisely the non-variable regions of the MHC I molecules." However, the new structure not only improves our understanding of the complex processes involved in loading MHC I molecules. It should also help select suitable candidates for vaccine development.
The research was published in Nature Communications.
More information: Ines Katharina Müller et al, Structure of an MHC I–tapasin–ERp57 editing complex defines chaperone promiscuity, Nature Communications (2022). DOI: 10.1038/s41467-022-32841-9
Journal information: Nature Communications
Provided by Goethe University Frankfurt am Main