An upgrade for magnetic resonance methods with a 1,000-fold amplifier

Researchers determine the structure and dynamics of proteins using NMR (Nuclear Magnetic Resonance) spectroscopy. Until now, however, much higher concentrations were necessary for in-vitro measurements of the biomolecules in solution than found in our body's cells. An NMR method enhanced by a very powerful amplifier, in combination with molecular dynamics simulation, now enables their detection and accurate characterization at physiological concentrations. This is reported by Dennis Kurzbach chemist at the University of Vienna and his colleagues in the journal Science Advances. The team demonstrated their new method with the example of a protein that influences cell proliferation and thus also potential tumor growth.
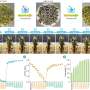
Currently, NMR spectroscopy is the only method that allows a complete description of the atomic structure of biomacromolecules in their native solution state. However, due to the inherently low sensitivity of the method, the samples must contain many more molecules per volume than physiologically common. To overcome this discrepancy, hyperpolarization (more precisely by Dissolution Dynamic Nuclear Polarization) can be used to achieve a 1,000-fold signal amplification in NMR measurements.
E-guitar vs. NMR—same principle
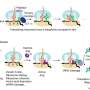
"Spectroscopy bears some similarities with an electric guitar: if the amplifier is too weak, you will hear very little if you do not hit the strings strongly," says Dennis Kurzbach of the Institute of Biological Chemistry, "meaning that you need a lot of material to see an NMR signal. With the new hyperpolarization amplifier you can now see something even at low concentration."
The researchers accomplished to measure biomolecules at concentrations as low as 1 micromole/liter (i.e. one-millionth of the usual concentration levels). The concentration thus approaches that of our cells. This is important because proteins can react to unnaturally high concentrations. They no longer do what they are supposed to do and suddenly behave differently.
Besides, a dissolution Dynamic Nuclear Polarization measurement typically provides one-dimensional spectra, which limits the obtained information. To describe proteins comprehensively under natural concentration conditions, the researchers employed molecular dynamics simulations: "We were also able to extrapolate the fingerprint we obtained of our molecule via NMR to its 'entire body,' i.e., its multidimensional structure," says Kurzbach.
Significant protein MAX described
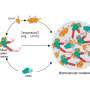
The value of this methodological advance is demonstrated using the ubiquitous transcription factor MAX. This protein can self-associate with various other proteins (i.e. protein dimerization). For example, MYC-MAX dimers have a great influence on the DNA copying processes in the cell.
With the new methods, MAX has been shown to adopt an undocumented conformation when concentrations approach physiological levels. "The folding spectrum of MAX is of crucial importance for working together with MYC and thus for the proliferation of healthy as well as diseased cells in the body," said ERC grantee Dennis Kurzbach, who is also deputy head of the Core Facility NMR at the Faculty of Chemistry.
The new method can help to better understand the process of cell proliferation to tumor growth and thus elucidate basic mechanisms for cancer development. This is just one of many potential fields of application for the new method—after all, thousands of proteins in our cells perform a wide variety of tasks, including digestion and regulation of DNA and RNA.
More information: Ludovica M. Epasto et al, Toward protein NMR at physiological concentrations by hyperpolarized water—Finding and mapping uncharted conformational spaces, Science Advances (2022). DOI: 10.1126/sciadv.abq5179
Journal information: Science Advances
Provided by University of Vienna