A fine-tuned gene editor that minimizes adverse consequences
The molecular tool CRISPR-Cas9 can be used to treat inherited blood disorders, but this may cause unintended genetic alterations. A team led by MDC researchers Klaus Rajewsky and Van Trung Chu has now presented an approach in Science Advances that minimizes such adverse consequences.
Hopes are high for the therapeutic potential of the CRISPR-Cas9 gene-editing tool. These "molecular scissors" can be used to very precisely cut out and repair gene mutations that are responsible for hereditary diseases. But despite the precision with which the tool is able to locate its target in the genome, its work is not yet completely error-free.
Sometimes, cuts are made at sites that are very similar to the target sequence, but are located in entirely different regions of the DNA. These mistakes, which scientists refer to as "off-target mutations," can have unexpected consequences. And even if CRISPR-Cas9 makes its cut at the correct site, errors can occur when the cut is repaired—a phenomenon known as "on-target mutations."
Spacing out the cuts
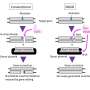
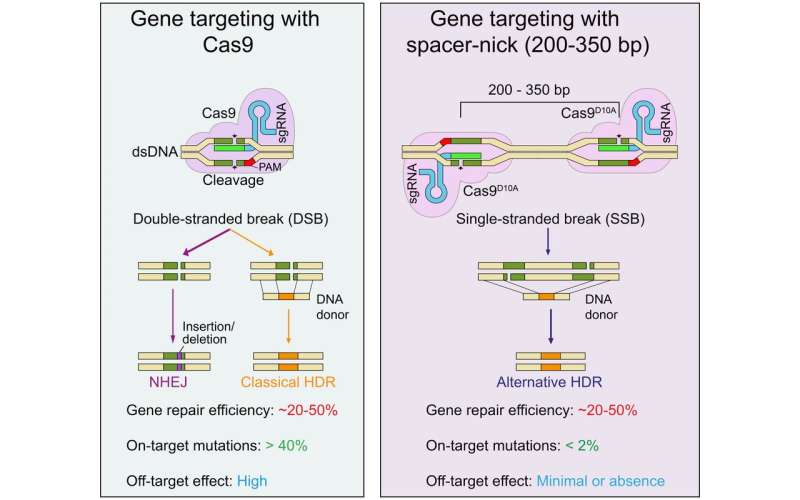
"These errors mainly occur because, in the classic method, both strands of the DNA molecule are cut at once," explains Professor Klaus Rajewsky, head of the Immune Regulation and Cancer Lab at the Berlin Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC). Together with other researchers from the MDC and the Humboldt-Universität zu Berlin, the scientist has now presented a new, refined approach in the journal Science Advances that has been dubbed "spacer-nick". This method employs a modified pair of molecular scissors, known as nickases, that make nicks on opposite strands of the DNA at two different points.
But the greater precision with which this gene-editing duo is able to detect and repair faulty genes is largely down to a spacer that the team has built into the tool. "We use this spacer to ensure that the two nicks are made 200 to 350 base pairs apart and that double-strand breaks in the DNA are avoided," explains Dr. Van Trung Chu, a scientist in the Rajewsky Lab and co-last author of the paper, along with Rajewsky himself. "Our experiments with hematopoietic stem cells and T cells have shown that this is the optimal distance for minimizing both on-target and off-target mutations," says Chu. "Any shorter, and we risk cutting through the entire DNA molecule—despite the use of two separate scissors."
Two other MDC teams—Professor Kathrin de la Rosa's Cancer & Immunology / Immune Mechanisms and Human Antibodies Lab, and Dr. Ralf Kühn's Genome Editing & Disease Models Lab, of which Chu is also a member—also made important contributions to the paper, especially in terms of detecting off-target mutations. "Spacer-nick is therefore also a good example of the successful collaboration between researchers working in the different labs at the MDC," says Rajewsky.
Effective and almost error-free
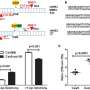
The scientists can even quantify the superiority of their fine-tuned genetic scissors and offset double nicks: "With the classic CRISPR-Cas9 method, on-target mutations occur in more than 40 percent of interventions," reports Chu. "The spacer-nick system can bring this down to under two percent." The success with off-target mutations, Chu explains, cannot be determined quite as easily and accurately: "All we can really say is that they occur relatively frequently when the classic genetic scissors are used, but have been a rare, if not non-existent, occurrence in our approach." What remains unclear for now is the exact mechanism by which the genetic material is repaired following the spacer-nick cuts. "It doesn't seem to happen via the well-known—and error-prone—NHEJ pathway," says Chu.
In terms of effectiveness, spacer-nick is on par with the conventional tool: "With both methods, we are able to successfully repair between 20 and 50 percent of the treated cells," says Chu. That is probably enough, he explains, to cure patients with an inherited blood disorder that stems from only a single altered gene. Examples of such disorders include beta thalassemia, which involves the faulty synthesis of the red blood pigment hemoglobin, or severe congenital neutropenia, which is characterized by a significantly reduced number of granulocytes—a type of white blood cell—and is associated with a severely weakened immune defense.
Fixing stem cells
Chu and Rajewsky hope that other researchers will take up their idea and test spacer-nick—first in animal models and then soon on the first human patients. Chu explains that the principle behind the therapy is simple: Blood-forming stem cells are taken from people with a monogenic inherited disorder using established methods. Spacer-nick then repairs the faulty genes directly in the cell culture. Once the genetic scissors have done their work, the repaired stem cells are administered back into the patient—where they produce new and, most importantly, healthy blood cells.
More information: Ngoc Tung Tran et al, Precise CRISPR-Cas–mediated gene repair with minimal off-target and unintended on-target mutations in human hematopoietic stem cells, Science Advances (2022). DOI: 10.1126/sciadv.abm9106
Journal information: Science Advances
Provided by Max Delbrück Center for Molecular Medicine