How bacteria store information to kill viruses (but not themselves)
During the last few years, CRISPR has grabbed headlines for helping treat patients with conditions as varied as blindness and sickle cell disease. However, long before humans co-opted CRISPR to fight genetic disorders, bacteria were using CRISPR as an immune system to fight off viruses.
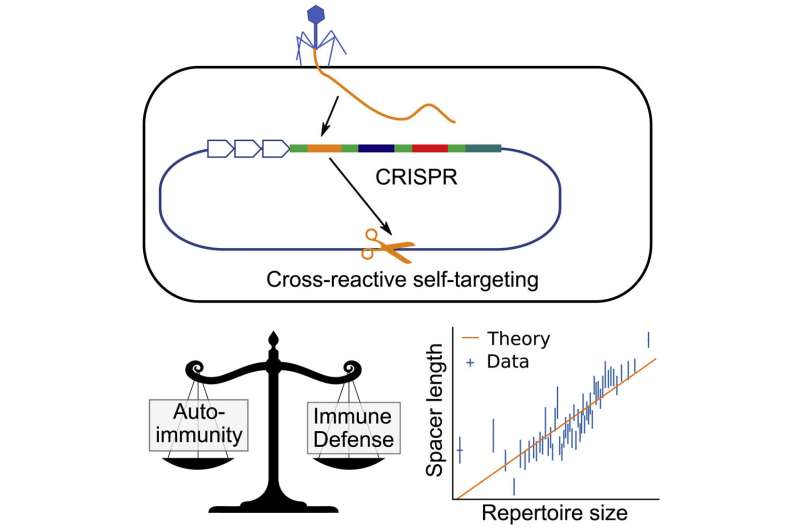
In bacteria, CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) works by stealing small pieces of DNA from infecting viruses and storing those chunks in the genes of the bacteria. These chunks of DNA, called spacers, are then copied to form little tags, which attach to proteins that float around until they find a matching piece of DNA. When they find a match, they recognize it as a virus and cut it up.
Now, a paper published in Current Biology by researchers from the University of Pennsylvania Department of Physics and Astronomy shows that the risk of autoimmunity plays a key role in shaping how CRISPR stores viral information, guiding how many spacers bacteria keep in their genes, and how long those spacers are.
Ideally, spacers should only match DNA belonging to the virus, but there is a small statistical chance that the spacer matches another chunk of DNA in the bacteria itself. That could spell death from an autoimmune response.
"The adaptive immune system in vertebrates can produce autoimmune disorders. They're very serious and dangerous, but people hadn't really considered that carefully for bacteria," says Vijay Balasubramanian, principal investigator for the paper and the Cathy and Marc Lasry Professor of Physics in the School of Arts & Sciences.
Balancing this risk can put the bacteria in something of an evolutionary bind. Having more spacers means they can store more information and fend off more types of viruses, but it also increases the likelihood that one of the spacers might match the DNA in the bacteria and trigger an autoimmune response.
Balasubramanian, along with coauthors Hanrong Chen of the Genome Institute of Singapore and Andreas Mayer of University College London, realized that the bacteria could get around this by having longer spacers. Similar to how a longer password might be harder to crack, a longer spacer would be less likely to match with the DNA of the bacteria itself. This means that bacteria with longer spacers would be able to have more spacers overall without the risk of triggering an autoimmune response.
With this idea in hand, the researchers built a mathematical model to calculate the ratio between spacer length and the total number of spacers that the bacteria should be able to store without risking an autoimmune response.
Once they worked out the mathematical model, they checked to see if their prediction held true in actual bacteria by looking at the CRISPR DNA of thousands of species and comparing the spacer length to the number of spacers stored.
The researchers found a consistent, tight relationship between spacer length and number of spacers.
"The surprise to me is that it matched so darn well just coming out of the box," says Balasubramanian. "This is a very simple theoretical framework. There's a risk of autoimmunity, but it's nice to have more immune memory, and you must balance these two considerations. It's just very, very rare that something so simple matches the data."
Balasubramanian says that the success of the model shows that this framework of simple, mathematical trade-offs might apply to more complex systems, such as the immune systems of vertebrates, including humans.
"Just by doing that statistical kind of reasoning you can make a lot of progress," he says. "So perhaps we can move back to vertebrate immunity and use the same techniques."
This study is also among the first to describe the importance of the autoimmune response in bacteria. Balasubramanian and his collaborators hope that future studies of CRISPR will consider the risk of autoimmunity.
As for future work in his group, he aims to explore how CRISPR stores information in response to evolving viruses. And while a statistical model of evolving bacterial genes might seem far removed from daily life, Balasubramanian says this work lays a foundation for a broader understanding of immunity, in ways that may enable deeper insight into viruses such as the seasonal flu or new SARS-CoV-2 variants.
Says Balasubramanian, "These are all pieces of a larger puzzle."
More information: Hanrong Chen et al, A scaling law in CRISPR repertoire sizes arises from the avoidance of autoimmunity, Current Biology (2022). DOI: 10.1016/j.cub.2022.05.021
Journal information: Current Biology
Provided by University of Pennsylvania