New technique may streamline the interpretation of genomes from less-studied animals
A new Cornell study, published in Nature Genetics, sheds light on a controversial debate in epigenetics—the set of molecular changes occurring on top of the genome that regulate how genes are turned on and off, but without changing a cell's DNA sequence.
The discovery may streamline the interpretation of new genomes from less-studied animals, researchers said.
Scientists studying epigenetics have long argued over the role of histone marks—little chemical tags hitched onto histones, the spool-like proteins that DNA wraps around inside the nucleus. Some researchers believe histone marks play a role in activating genes and kicking off transcription—the process of copying DNA into RNA to make proteins or for other cellular functions. The new study, however, finds that histone marks are laid down during the transcription process and don't play a regulatory role.
"This potentially has big implications for the way that we interpret genomes," said Charles Danko, the Robert N. Noyce Associate Professor in Life Science and Technology at the Baker Institute in the College of Veterinary Medicine and corresponding author of the study. "We would like to apply these same technologies to interpret the genome function in horse, dog and other veterinary species where there is not as much funding for research."
Zhong Wang, formerly of the Danko lab, and now a professor at Dalian University in China, is a co-lead author on the study, "Prediction of Histone Post-Translational Modification Patterns Based on Nascent Transcription Data," which published March 10 in Nature Genetics.
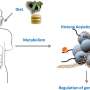
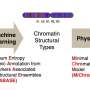
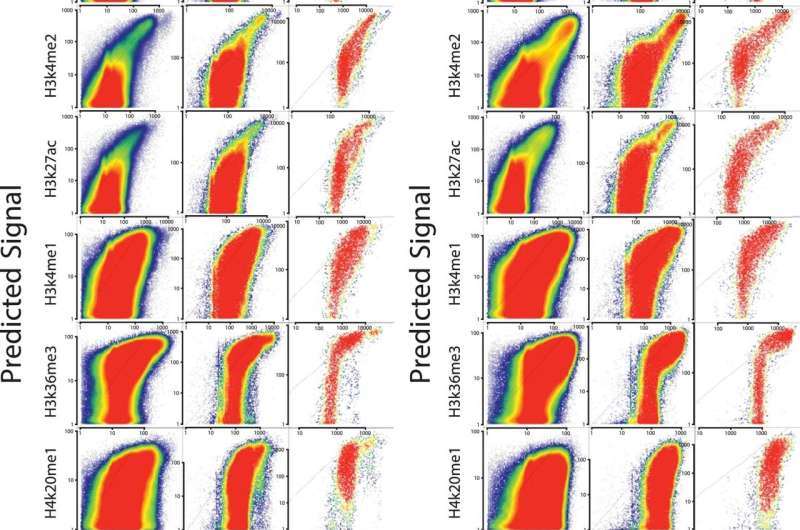
Wang used publicly available data sets of transcription in mice and humans to develop a computer program that predicts where histone marks occur in the genome. The program, called dHIT, takes data from a single experiment showing where transcription is actively occurring on the chromosomes. Researchers can use a technique called ChRO-seq, which was developed in the Danko lab, or a related technique called PRO-seq, developed at Cornell, to generate the transcription data.
Danko's group used this approach successfully to predict histone locations in mouse, human and horse cells. They showed dHIT works almost as well as biological experiments that directly measure the location of each mark.
Dr. Doug Antczak, the Dorothy Havemeyer McConville Professor of Equine Medicine at Baker, and colleagues from the Functional Annotation of Animal Genomes (FAANG) project contributed data and samples from horses for these experiments.
Alexandra Chivu, a doctoral student in the Danko lab and co-lead author, used dHIT in combination with experimental techniques measuring histone marks to find out what happens when transcription is blocked in a cell. She showed that without transcription, histone marks disappear, confirming that their intricate patterns often depend on transcription.
Before the development of dHIT, entire scientific consortia of researchers would work together on a single genome from a model organism to identify the location of histone marks. These new tools greatly simplify that process.
Next, Danko plans to tease out the actual role of histone marks by removing them and measuring the impact on transcription. "If we accept that these marks are parts of the machinery of transcription, then a really interesting question is, what parts of that machinery are they involved in?" he said. "What is their role if it's not regulation?"
More information: Zhong Wang et al, Prediction of histone post-translational modification patterns based on nascent transcription data, Nature Genetics (2022). DOI: 10.1038/s41588-022-01026-x
Journal information: Nature Genetics
Provided by Cornell University