High-resolution mapping method for G4 DNA structures developed
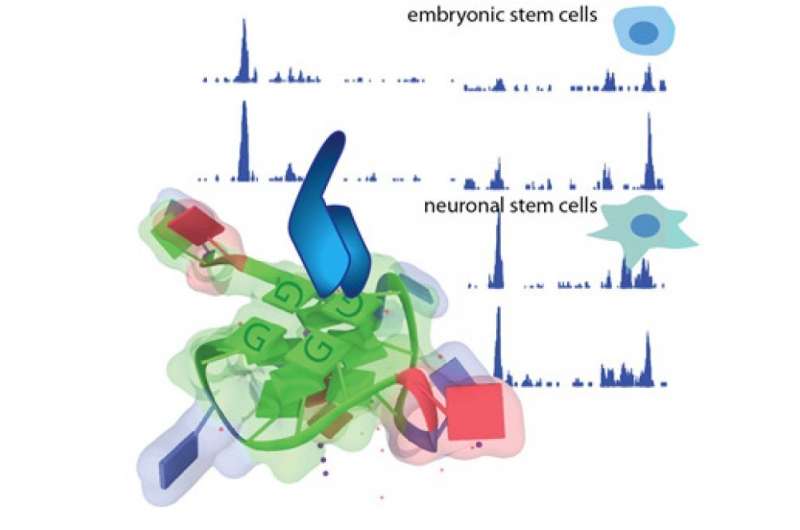
While investigating the unusual G quadruplex DNA structure (G4), the Simon Elsässer group has developed a more accurate method for mapping these structures in the genome. G4 CUT&Tag revealed numerous G4s in the human and mouse genome that were previously not detected. The study has been published in Nucleic Acid Research. These findings may be used for drug development for cancer therapy and provide an important tool to evaluate drug action and safety.
When the canonical double-stranded helix of DNA is melted, the individual strands can adopt alternative folds. In so-called G quadruplex structure, four guanine bases arrange in a planar quartet instead of pairing with cytosine in the usual manner. Several G quartets stacked on top of each other form the G quadruplex structure (G4). G4s were first spotted 20 years ago at the ends of chromosomes in so-called telomeric repeat DNA. But by now, researchers have uncovered that G4s come in many different variations and appear frequently also in other regions of the human genome.
G4s location, formation, and function yet unknown
Where exactly G4s are located, how they form, and what they could be good for remains somewhat of a mystery to researchers. In Simon Elsässer's group, doctoral student Jing Lyu has developed a new method to map G4 structures genome-wide at high resolution.
"While genome-wide mapping methods for G4 were developed earlier, we were concerned that available methodologies did not retain the delicate G4 structures through the protocol and hence only allowed identification of the most abundant and stable G4s in the genome. Applying an in-situ approach, in which antibodies against the G4 structures are diffused into lightly permeabilized cells, we were able to 'mark' the DNA sequences that formed a G4," says Jing Lyu, Ph.D. student in Simon Elsässer's group at the Department of Medical Biochemistry and Biophysics.
More accurate CUT&Tag method for G4
This method, known as CUT&Tag, relies on a transposase enzyme that is recruited via the antibody and inserts an oligonucleotide into the genomic sequence that can later be used to amplify and sequence the targeted sites. The resulting G4 CUT&Tag method provided Lyu with a much better signal-to-noise ratio than existing methods and revealed numerous G4s in the human and mouse genome that were previously not detected.
Mapping G4s for the first time in mouse embryonic stem cells, Lyu discovered that, in addition to most active gene promoters, enhancers bound by pluripotency factors also featured G4s. The ability to form G4s seemed to make a clearcut distinction between enhancers active during pluripotency and tissue-specific enhancers activated during differentiation.
Implications for cancer therapy and drug action and safety
"In the future we would like to investigate if and how enhancer G4s play a role in regulating gene activity and 3D genome organization" says Simon Elsässer, researcher and principal investigator at the Department of Medical Biochemistry and Biophysics.
Small molecules stabilizing G4 structures receive strong interest in drug development for cancer therapy, and genome-wide mapping of G4s will provide an important tool to evaluate drug action and safety.
The study, available as a preprint on bioRxiv since April, is now published in Nucleic Acid Research.
More information: Jing Lyu et al, Genome-wide mapping of G-quadruplex structures with CUT&Tag, Nucleic Acids Research (2021). DOI: 10.1093/nar/gkab1073
Journal information: Nucleic Acid Research , Nucleic Acids Research
Provided by Karolinska Institutet