Mammal biodiversity shapes disease risk
To predict the spillover of pathogens from wildlife to humans and limit future pandemics, it is important to understand where wildlife disease risk is highest. A team of researchers, led by Wageningen University & Research (WUR) in The Netherlands, report that mammal biodiversity is an important mechanism driving wildlife disease risk. They identified hotspots of disease risk using the predicted distributions and abundances of more than 4,000 mammal species worldwide.
Mammals can carry pathogens that are transmittable to humans (so-called zoonotic pathogens). SARS-CoV-2, the virus that causes COVID-19, is one example. The researchers found that spatial variation in mammal assemblage richness and abundance shapes wildlife disease risk. "With one quarter of all mammal species threatened by extinction, we urgently need to understand how biodiversity loss influences zoonotic disease risk," says Yingying Wang. Wang initiated this study as part of her Ph.D. at WUR. She is currently affiliated with University of Jyväskylä in Finland.
She and her colleagues first used information on climate and the habitat preferences, body masses, and diets of mammals, to model terrestrial mammal assemblages worldwide. That is: how many individuals of what species live together and where? Based on these assemblages, they were able to calculate disease risk using an ecological version of the epidemiological parameter R-naught. This "community-level" R-naught represents the basic reproduction number for pathogens that infect multiple hosts. This approach was then extended to forecast disease risk.
Hotspots
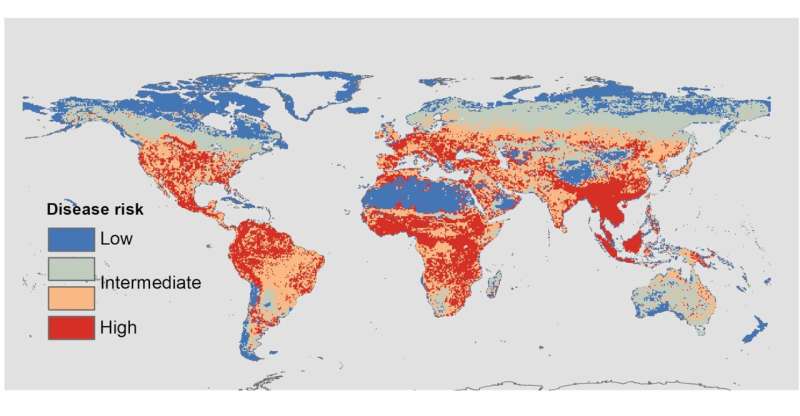
The results predicted high disease risk in tropical areas where biodiversity is highest, but there were also risky temperate areas. Risk of density-dependent diseases was high in the equatorial tropics and in temperature Europe, North America, and Asia, while risk of frequency-dependent diseases was high in the northernmost parts of North America and Eurasia and in Oceania.
These predictions fit well with past global patterns of disease outbreaks. However, there were still some surprises according to Kevin Matson, wildlife ecologist at WUR. Matson: "People often view the tropics as risky in terms of the emergence of new diseases from wildlife. But our predictions highlight the possibility that new outbreaks may also occur outside the tropics." According to Matson, one reason may be that some mammals, in particular rodents, in temperate areas are better at sharing some viruses among host species, though he notes that this possibility was not explicitly tested in the current research.
Biodiversity loss and disease risk
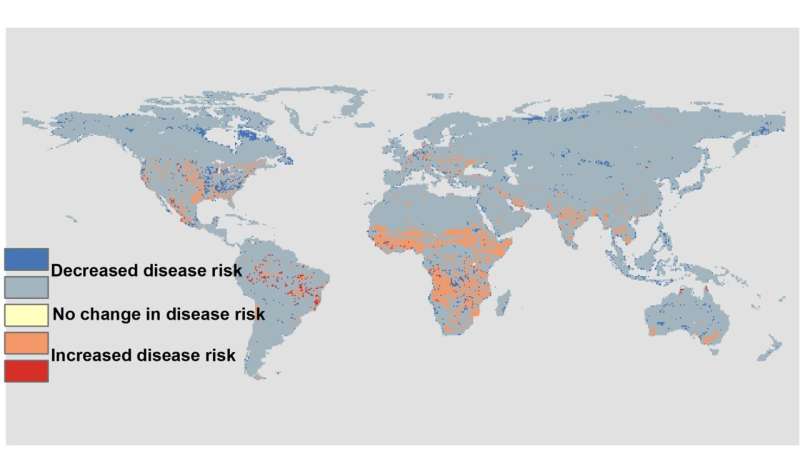
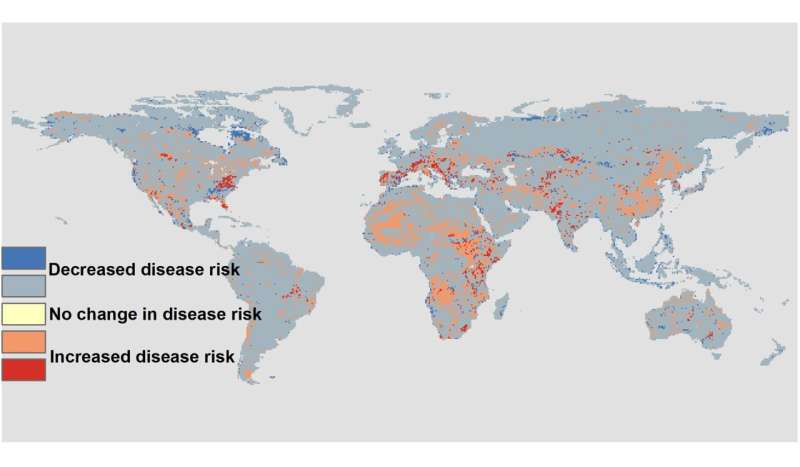
In the face on an ongoing biodiversity crisis caused by habitat loss, climate change, and other global forces, the researchers also wondered how wildlife disease risk might change in the coming decades. To do this, they extended their model to incorporate environmental changes anticipated under two different pathways: one with a relatively rosy view of the future and a second, more pessimistic one. It turns out that species do not have to go locally extinct to impact disease risk, as often assumed. Substantial changes in disease risk can occur without extinction of any species, instead being influenced by changes in relative abundance.
"A little more of this species and a little less of that one can do the trick," says Fred de Boer, wildlife ecologist and colleague of Matson at WUR. "For example, large species may regulate the abundance of smaller ones, which means we have to consider that conserving larger species could serve as an important control measure. Findings as such this one force us to acknowledge the nuance and complexity of relationships between biodiversity and disease. It is exciting that our predictions fit so well with past global patterns of disease outbreaks. This suggests that we've made a step in the right direction of understanding how the relationship between disease risk and biodiversity really works," De Boer concludes.
The study is published in the journal Global Change Biology.
More information: Yingying X. G. Wang et al, Mammal assemblage composition predicts global patterns in emerging infectious disease risk, Global Change Biology (2021). DOI: 10.1111/gcb.15784
Journal information: Global Change Biology
Provided by Wageningen University