AI used to predict unknown links between viruses and mammals
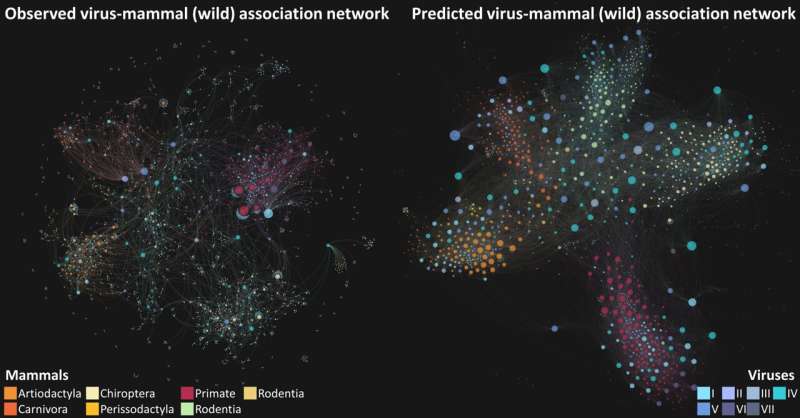
A new University of Liverpool study could help scientists mitigate the future spread of zoonotic and livestock diseases caused by existing viruses.
Researchers have used a form or artificial intelligence (AI) called machine-learning to predict more than 20,000 unknown associations between known viruses and susceptible mammalian species. The findings, which are published in Nature Communications, could be used to help target disease surveillance programmes.
Thousands of viruses are known to affect mammals, with recent estimates indicating that less than 1% of mammalian viral diversity has been discovered to date. Some of these viruses such as human and feline immunodeficiency viruses have a very narrow host range, whereas others such as rabies and West Nile viruses have very wide host ranges.
"Host range is an important predictor of whether a virus is zoonotic and therefore poses a risk to humans. Most recently, SARS-CoV-2 has been found to have a relatively broad host range which may have facilitated its spill-over to humans. However, our knowledge of the host range of most viruses remains limited," explains lead researcher Dr. Maya Wardeh from the University's Institute of Infection, Veterinary and Ecological Sciences.
To address this knowledge gap, the researchers developed a novel machine learning framework to predict unknown associations between known viruses and susceptible mammalian species by consolidating three distinct perspectives—that of each virus, each mammal, and the network connecting them, respectively.
Their results suggests that there are more than five times as many associations between known zoonotic viruses and wild and semi-domesticated mammals than previously thought. In particular, bats and rodents, which have been associated with recent outbreaks of emerging viruses such as coronaviruses and hantaviruses, were linked with increased risk of zoonotic viruses.
The model also predicts a five-fold increase in associations between wild and semi-domesticated mammals and viruses of economically important domestic species such as livestock and pets.
Dr. Wardeh said: "As viruses continue to move across the globe, our model provides a powerful way to assess potential hosts they have yet to encounter. Having this foresight could help to identify and mitigate zoonotic and animal-disease risks, such as spill-over from animal reservoirs into human populations."
Dr. Wardeh is currently expanding the approach to predict the ability of ticks and insects to transmit viruses to birds and mammals, which will enable prioritisation of laboratory-based vector-competence studies worldwide to help mitigate future outbreaks of vector-borne diseases.
More information: Maya Wardeh et al, Divide-and-conquer: machine-learning integrates mammalian and viral traits with network features to predict virus-mammal associations, Nature Communications (2021). DOI: 10.1038/s41467-021-24085-w
Journal information: Nature Communications
Provided by University of Liverpool