Universal mechanism of regulation in plant cells discovered
All plant cells obtain their energy mainly from two organelles they contain—chloroplasts (responsible for photosynthesis) and mitochondria (responsible for the biochemical cycle of respiration that converts sugars into energy). However, a large number of a plant cell's genes in its mitochondria and chloroplasts can develop defects, jeopardizing their function. Nevertheless, plant cells evolved an amazing tool called the RNA editosome (a large protein complex) to repair these kinds of errors. It can modify defective messenger RNA that result from defective DNA by transforming (deamination) of certain mRNA nucleotides.
Automatic error correction in plant cells
Automatic error correction in plants was discovered about 30 years ago by a team headed by plant physiologist Axel Brennicke and two other groups simultaneously. This mechanism converts certain cytidine nucleotides in the messenger RNA into uridine in order to correct errors in the chloroplast DNA or mitochondrial DNA. RNA editing is therefore essential to processes such as photosynthesis and cellular respiration in plants. Years later, further studies showed that a group of proteins referred to as PPR proteins with DYW domains play a central role in plant RNA editing. These PPR proteins with DYW domains are transcribed in the cell nucleus and migrate through the cells to chloroplasts and mitochondria. However, they are inactive on their way to these organelles. Only once they are within the organelles do they become active and execute their function at a specific mRNA site. How this activation works, however, has been a mystery until now.
It doesn't work in a test tube
For many years, it was not possible to synthetically produce these DYW-type PPR proteins in the laboratory to study their function and structure more closely. Only now has a German-Japanese team headed by structural biologist and biochemist Dr. Gert Weber from the Joint Protein Crystallography Group at Helmholtz-Zentrum Berlin and Freie Universität Berlin succeeded in doing so.
Now: 3D structure of the key protein decoded
Prof. Mizuki Takenaka's group had previously been able to produce the DYW domain in bacteria. Takenaka has been conducting research at Kyoto University since 2018 and previously worked in Axel Brennicke's laboratory in Ulm, Germany. Tatiana Barthel (University of Greifswald and now at HZB) was then able to grow the first protein crystals of the DYW domain. A large number of these delicate crystals have now been analyzed at the MX beamlines of BESSY II so that the three-dimensional architecture of the DYW domain could be decoded. "Thanks to the Joint Research Group co-located at HZB and FU Berlin, we have the capability of beam time for measurements very quickly when needed, which was crucial," says Dr. Manfred Weiss, who is responsible for the MX beamlines at BESSY II and co-author of the study.
Mechanism of activation discovered
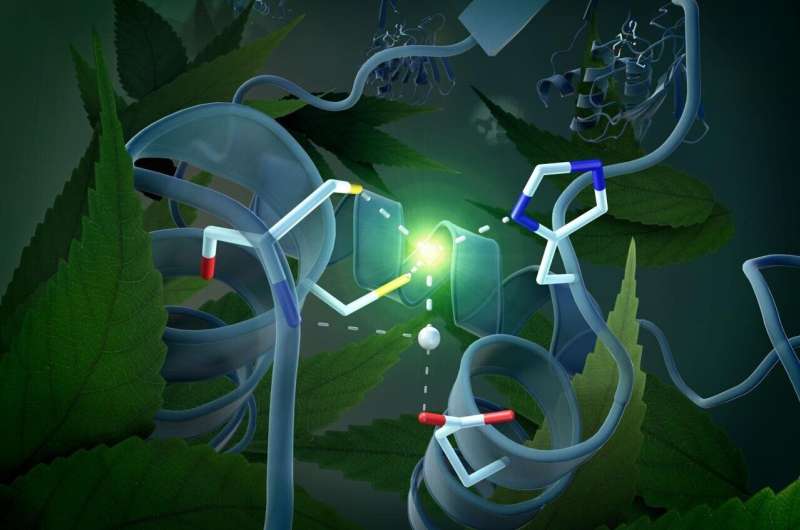
This three-dimensional architecture has actually provided the crucial clue to the mechanism of DYW domain activation that applies to all plants. It is due to a zinc atom located in the center of the DYW domain that can accelerate the deamination of cytidine to uridine like a catalyst. For this to happen, however, the zinc must be optimally positioned. The activation switch is provided by a very unusual gating domain in the immediate vicinity of the catalytic center—the structural analysis shows that this gating domain can assume two different positions, thereby switching the enzyme on or off. "The movement of the gating domain regulates the extent to which the zinc ion is available for the catalytic reaction," Weber explains.
A molecule like scissors
Now it has become clear why getting DYW-type PPR proteins to react with RNA in the test tube has been difficult until now: These PPR proteins are nominally inactive and require activation. In the plant cells, they are first produced in the cell nucleus and then very likely migrate in an inactivated state to the organelles, where they become activated. "This is ideal, because otherwise these molecules would be active along the way, altering various RNA molecules in an uncontrolled fashion harmful to the cell," says Weber.
Universal repair tool
This work is a breakthrough for plant molecular biology because it describes an additional level of sophisticated regulation in chloroplasts and mitochondria. The results are fundamental for plant science, but they could also play a role in our daily lives someday. The DYW domain might provide a useful tool for controllable and site-specific C-to-U and U-to-C RNA editing. This could open up new bioengineering and medical applications, such as reprogramming certain mitochondrial genes without changing a cell's nuclear DNA.
More information: Mizuki Takenaka et al, DYW domain structures imply an unusual regulation principle in plant organellar RNA editing catalysis, Nature Catalysis (2021). DOI: 10.1038/s41929-021-00633-x
Journal information: Nature Catalysis
Provided by Helmholtz Association of German Research Centres