Genetic treasure trove for malaria researchers
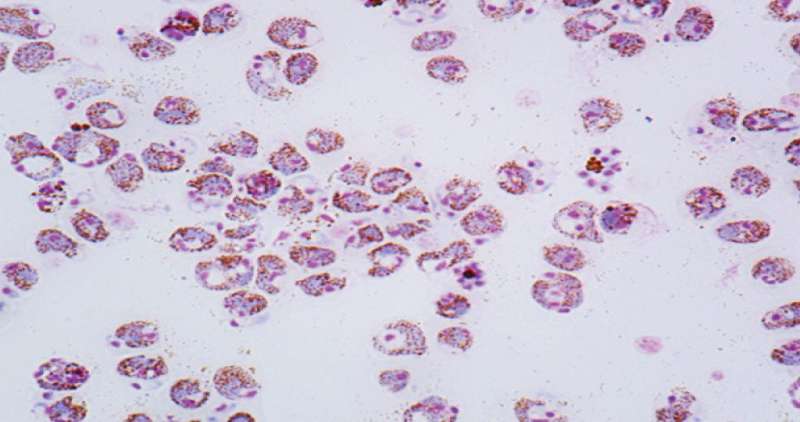
Detailed and extensive genome sequencing of a subspecies of rat-infecting malaria parasites should instruct human malaria research.
A new extensive genetic resource of rat-infecting malaria parasites may help advance the development of malaria prevention and treatment strategies. This trove of genome and phenome information has been published by a team of KAUST researchers, along with colleagues in Japan, and the datasets have been made publicly available for malaria researchers.
Rodent malaria parasites are closely related to human parasites, but are easier to study because they can be grown in laboratory mice. "Investigations on rodent malaria parasites have played a key role in revealing many aspects of fascinating biology across their life-cycle stages," says KAUST bioscientist Arnab Pain, who led the sequencing effort, in collaboration with Richard Culleton of Nagasaki University's Institute of Tropical Medicine.
Most research on these parasites to date has involved three specific species, but a fourth, called Plasmodium vinckei, hasn't received much attention.
Pain, Culleton and the team generated a comprehensive genetic resource for this species and also sequenced genomes of seven isolates belonging to two of the other species, P. yoelii and P. chabaudi.
Sequencing the genomes of ten isolates from five subspecies of P. vinckei from tropical Africa revealed that they have widely diverged from their common ancestor. The evolutionary pressures on each of the subspecies varies greatly according to the regions where they are mainly found.
The sequencing efforts clarified aspects of the evolutionary tree of rat malaria parasites and also led to the naming of three new subspecies: P. yoelii cameronensis, P. chabaudi esekanensis and P. vinckei baforti.
The research describes in detail genetic and phenotypic variations between the subspecies, which is likely to help studies that aim to understand the functions of malaria parasite genes.
The scientists were also able to genetically modify a subspecies of P. vinckei to carry a fluorescent protein. This demonstrates that investigations on gene function that involve modifying or removing a target gene can be conducted in this subspecies.
"We hope our resource will provide the research community with a diverse set of parasite models to play with. This resource can be put to use to identify genes that influence the malaria parasite's virulence, drug resistance and transmissibility in mosquitoes," says Abhinay Ramaprasad, the first author of this study, which he conducted during his Ph.D. at KAUST.
The resource has been well received by the research community: "What a rich set of resources," wrote Jane Carlton, Director of New York University's Center for Genomics & Systems Biology. "The development of a model system once took decades, but with the aid of next-generation sequencing … and enhanced molecular biology techniques, Ramaprasad et al. have fast-tracked the establishment of P. vinckei as a useful additional experimental model for malaria."
More information: Abhinay Ramaprasad et al, Plasmodium vinckei genomes provide insights into the pan-genome and evolution of rodent malaria parasites, BMC Biology (2021). DOI: 10.1186/s12915-021-00995-5
Jane M. Carlton, A cornucopia of research resources for the fourth rodent malaria parasite species, BMC Biology (2021). DOI: 10.1186/s12915-021-01019-y
Journal information: BMC Biology