Selective mRNA degradation via autophagy: A novel role for autophagy in gene regulation
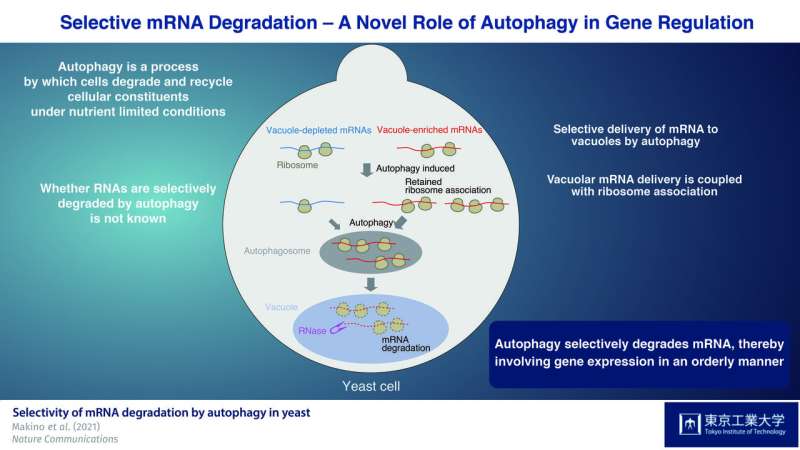
Optimal cell function requires a fine balance between the synthesis and degradation of biomolecules. Autophagy is the process by which cells degrade and recycle their own components, helping to clean up and maintain the cell's internal environment and ensure the smooth functioning of cellular processes. Autophagy is strongly induced when cells are subjected to stresses like nutrient deprivation, acting under such conditions to supply nutrients through its breakdown of unneeded cellular material.
Autophagy substrates are delivered to vacuoles in yeast or lysosomes in mammals for degradation by double-membrane vesicles called "autophagosomes." While autophagy was originally considered a non-selective process that isolates substrates in the cytoplasm of the cell in a random manner, studies have reported that certain cellular components, such as a subset of proteins and damaged or superfluous cell organelles, are isolated in a selective manner. In contrast to this well-established targeting of organelles and proteins by autophagy, the question of whether RNAs are subjected to autophagy and if they are selectively degraded has remained unanswered.
In their latest study, which was published in Nature Communications, researchers from the Tokyo Tech and RIKEN conducted a detailed analysis of the preferential degradation by autophagy of messenger RNAs (mRNAs), which contain the information required to make cellular protein and bind ribosomes for protein synthesis. Corresponding author Prof. Yoshinori Ohsumi of the Tokyo Tech, who was awarded the 2016 Nobel Prize in Physiology or Medicine for his pioneering work in the field of autophagy, explained the group's findings, stating "We have previously shown that RNA delivered to the vacuole via autophagy in yeast cells, where it is degraded by vacuolar nucleases. The question of whether RNA degradation by autophagy occurs preferentially, however, remains unaddressed. This difficult to address question was the starting point of this project."
As RNAs that accumulate in the vacuole are enzymatically degraded by the nuclease Rny1, they first constructed a yeast strain lacking this enzyme. Using this strain, they were able to isolate and identify RNAs that accumulated in the vacuole. Next, they used the drug rapamycin, which is known to induce autophagy, to assess unique features of mRNA species delivered to the vacuole in Rny1-deficient cells when autophagy is induced. Critically, they discovered that autophagy-mediated mRNA delivery to vacuoles is selective, not random, in nature.
The researchers then characterized the different mRNA species by conducting a broad analysis of the types of mRNAs in these cells, identifying 'vacuole-enriched' and 'vacuole-depleted' mRNAs. Interestingly, housekeeping mRNAs, such as those encoding proteins involved in amino acid biosynthesis, were most likely to be delivered to vacuoles. In contrast, mRNAs required for the synthesis of proteins with regulatory functions, such as protein kinases, were predominantly detected in the vacuole-depleted mRNA fraction.
Furthermore, they demonstrated that mRNAs undergoing translation are delivered to the vacuole, which is suggested to be a translation-dependent process. Moreover, persistent ribosome-mRNA association upon rapamycin treatment was found to be a key determinant of vacuolar mRNA delivery during autophagy-mediated degradation.
Dr. Makino and Prof. Ohsumi highlighted the importance of autophagy in gene regulation, remarking, "Our findings suggest that autophagy regulates mRNA degradation at the translation step, thereby enabling a rapid and sensitive switch from ribosome-associated mRNAs to expression of mRNAs that are essential for an effective response to stress. Preferential degradation of ribosome-mRNAs by autophagy is therefore very likely to determine the fate of individual mRNAs as cells adapt to new conditions."
More information: Shiho Makino et al. Selectivity of mRNA degradation by autophagy in yeast, Nature Communications (2021). DOI: 10.1038/s41467-021-22574-6
Journal information: Nature Communications
Provided by Tokyo Institute of Technology