Synthesis of potent antibiotic follows unusual chemical pathway
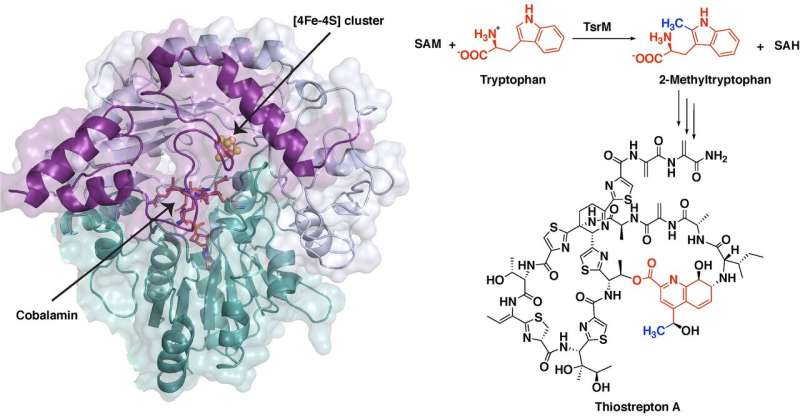
Images of a protein involved in creating a potent antibiotic reveal the unusual first steps of the antibiotic's synthesis. The improved understanding of the chemistry behind this process, detailed in a new study led by Penn State chemists, could allow researchers to adapt this and similar compounds for use in human medicine.
"The antibiotic thiostrepton is very potent against Gram-positive pathogens and can even target certain breast cancer cells in culture," said Squire Booker, a biochemist at Penn State and investigator with the Howard Hughes Medical Institute. "While it has been used topically in veterinary medicine, so far it has been ineffective in humans because it is poorly absorbed. We studied the first steps in thiostrepton's biosynthesis in hopes of eventually being able to hijack certain processes and make analogs of the molecule that might have better medicinal properties. Importantly, this reaction is found in the biosynthesis of numerous other antibiotics, and so the work has the potential to be far reaching."
The first step in thiostrepton's synthesis involves a process called methylation. A molecular tag called methyl group, which is important in many biological processes, is added to a molecule of tryptophan, the reaction's substrate. One of the major systems for methylating compounds that are not particularly reactive, like tryptophan, involves a class of enzymes called radical SAM proteins.
"Radical SAM proteins usually use an iron-sulfur cluster to cleave a molecule called S-adenosyl-L-methionine (SAM), producing a "free radical" or an unpaired electron that helps move the reaction forward," said Hayley Knox, a graduate student in chemistry at Penn State and first author of the paper. "The one exception that we know about so far is the protein involved in the biosynthesis of thiostrepton, called TsrM. We wanted to understand why TsrM doesn't do radical chemistry, so we used an imaging technique called X-ray crystallography to investigate its structure at several stages throughout its reaction."
In all radical SAM proteins characterized to date, SAM binds directly to the iron-sulfur cluster, which helps to fragment the molecule to produce the free radical. However, the researchers found that the site where SAM would typically bind is blocked in TsrM.
"This is completely different from any other radical SAM protein," said Booker. "Instead, the portion of SAM that binds to the cluster associates with the tryptophan substrate and plays a key role in the reaction, in what is called substrate-assisted catalysis."
The researchers present their results in an article appearing Jan. 18 in the journal Nature Chemistry.
In solving the structure, the researchers were able to infer the chemical steps during the first part of thiostrepton's biosynthesis, when tryptophan is methylated. In short, the methyl group from SAM transfers to a part of TsrM called cobalamin. Then, with the help of an additional SAM molecule, the methyl group transfers to tryptophan, regenerating free cobalamin and producing the methylated substrate, which is required for the next steps in synthesizing the antibiotic.
"Cobalamin is the strongest nucleophile in nature, which means it is highly reactive," said Knox. "But the substrate tryptophan is weakly nucleophilic, so a big question is how cobalamin could ever be displaced. We found that an arginine residue sits under the cobalamin and destabilizes the methyl-cobalamin, allowing tryptophan to displace cobalamin and become methylated."
Next the researchers plan to study other cobalamin-dependent radical SAM proteins to see if they operate in similar ways. Ultimately, they hope to find or create analogs of thiostrepton that can be used in human medicine.
"TsrM is clearly unique in terms of known cobalamin-dependent radical SAM proteins and radical SAM proteins in general," said Booker. "But there are hundreds of thousands of unique sequences of radical SAM enzymes, and we still don't know what most of them do. As we continue to study these proteins, we may be in store for many more surprises."
More information: Structural basis for non-radical catalysis by TsrM, a radical SAM methylase, Nature Chemistry (2021). DOI: 10.1038/s41589-020-00717-y , www.nature.com/articles/s41589-020-00717-y
Journal information: Nature Chemistry
Provided by Pennsylvania State University