Building with DNA
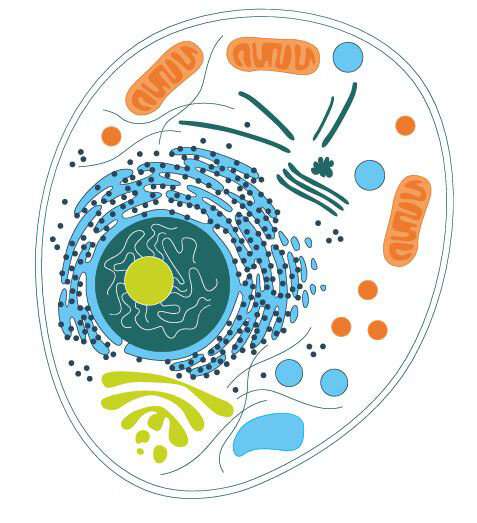
Life on Earth developed from inanimate components. Can we recreate this process in the laboratory, and what tools do we need for this? Using DNA origami, the art of folding at a scale of just a few millionths of a millimetre, we are able to reconstruct individual cellular components. They may be capable of taking over important tasks in our bodies in future.
A living cell consists of separate yet connected functional units: a cell envelope, organelles, metabolism and the genome. "What I cannot create, I cannot understand"—this statement by the physicist and Nobel laureate Richard Feynman holds true for living cells. It is still impossible to create an artificial cell from scratch. But history teaches us that what was unimaginable yesterday and may become reality tomorrow: at the beginning of the 19th century, for example, most chemists were convinced that urea could not be created artificially from inanimate matter. They believed that a "vital spark" was required to produce an organic substance such as this component of urine. In 1828, the chemist Friedrich Wöhler then manufactured urea from ammonium chloride and silver cyanate.
For the first time, humans were able to create a substance in a test tube which had previously only been known from living organisms. Today, synthetic biology may stand at the point where organic chemistry was before artificial urea synthesis. But a cell is far more complex than the chemistry of ammonium salts.
Therefore, the recipe for success in synthetic biology is simplification: Researchers select only the most important components and put them together in cell-like compartments. In a so-called bottom-up approach, they create minimal modules, each of which copies one specific function of a living cell: One module can, for instance, convert light into chemical energy, another one responds to stimuli, while a third enables movement. But putting together the modules to form a complete functional synthetic cell is still very difficult because it is not always clear how exactly they fit together. In addition, their interactions are extremely complex.
Artifical cells should be programmable and independent
Perhaps it would be more straight forward to change perspectives. Instead of combining existing modules to reconstruct the original as accurately as possible, researchers could use new tools and materials. This leaves room for creative solutions. My team and I are applying such a "de novo" approach to build an artificial cell from human-made components. Instead of copying life as we know it, we aim for a unique original. Our artificial cell should be programmable and act independently—like a miniature robot, as a link between the animate and the inanimate world. But what tools and what materials are suitable for building such synthetic components? In any case, they have to be programmable precision tools, which can provide a high number of copies of molecular units, precisely engineered for various different functions.
Letter sequence determines the form of DNA molecules
In our opinion, one suitable tool is DNA origami, the art of folding with DNA. In this approach, DNA is not used for hereditary information storage like in nature, but as a building material. The spiral-shaped DNA double helix is unwound and split into individual strands. One long individual strand can now be folded into a desired shape using many short, artificially created DNA sequences. We calculate the required DNA sequences based on a 3-D drawing and then combine them in a test tube. This process creates trillions of identical copies of the previously designed shape in a drop of water, each only a few millionths of a millimetre in size. It may sound like magic, but it's simply physics: due to the prescribed base pairings, the DNA strands self-assemble to maximise their fit and create the designed three-dimensional structure, which can take over a specific task in a cell. In this way, we have produced artificial membrane pores from DNA—components which are difficult to isolate from cells. But it does not always have to be complicated structures. Even a single, chemically modified DNA double helix is sufficient to connect two components in a cell. Nature often uses hundreds of linker proteins—for example, to tie the cytoskeleton to the cell membrane.
It appears highly challenging to isolate them individually and integrate them into synthetic cells. That's why we have chosen to take a short cut and use DNA as an artificial link. This link can be broken and reformed based on external stimuli such as temperature changes. At the end, we need to combine the different components inside a compartment in order to mimic a cell.
We are already able to divide such cell-like compartments. Yet assembling it in the first place is not easy, because the cell envelope is particularly fragile. Once the membrane, which consists of a thin fatty acid layer, has been formed, the insertion of components becomes challenging.
A boundary layer of droplets as artificial cell envelope
That's why we have developed a method which superficially resembles cocktail shaking: we first layer the components on top of each other in a test tube and then shake them to create a droplet emulsion. An artificial cell envelope forms at the boundary layer of the droplets and encapsulate the components. In this simple way, we can include many different components. Also microfluidics and 3-D printing are helpful tools. With them at hand, we can dedicate ourselves to the next challenge: the development of an information storage system for synthetic cells. The emergence of life on Earth took billions of years. When will the laboratory experiment be successful? Humans could be much quicker because, instead of waiting for a series of lucky coincidences like in natural evolution, we pursue clear goals with synthetic biology. This gives us hope that an artificial living model system may soon become a reality. Then, at the latest, an ancient question will gain new meaning: What is life, and could it perhaps look very different?
Provided by Max Planck Society