Scientists visualize the structure of a key enzyme that makes triglycerides
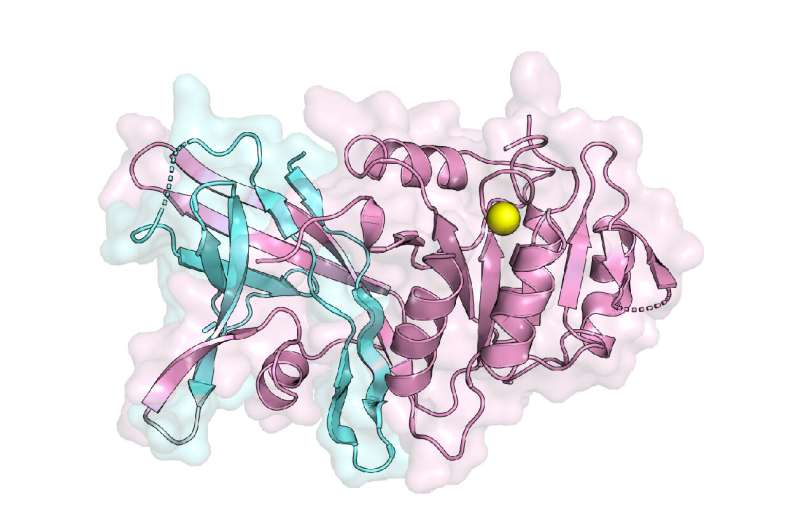
The first structure of a lipin enzyme, which carries out an important step in the production of triglycerides, the main reservoir for long-term energy storage, will help scientists to better understand how lipins regulate the production of triglycerides. Led by Mike Airola, Ph.D., of the Department of Biochemistry and Cell Biology in the College of Arts and Sciences and Renaissance School of Medicine at Stony Brook University, the structure also provides scientists with insights as to why mutations in the enzyme cause a loss of activity that leads to abnormal production of triglycerides implicated in heart disease, obesity and diabetes. The study will published in Nature Communications.
Lipins complete the second to last step of triglyceride production. But when mutations interrupt lipin functions, the body loses its ability to store fat properly, thus potentially triggering a wide variety of metabolic-related conditions. Scientists have unsuccessfully tried to create the first visual structure a lipin enzyme since these enzymes were identified in 2001. Now Airola and colleagues successfully explain in the paper their first crystal structure of a specific enzyme called lipin PAP.
"This structure answers a long-standing question for how two essential regions, N-lip and C-lip, which are located on opposite ends of this protein in humans, come together to form a function enzyme to help make triglycerides," explains Airola, Assistant Professor in the Department of Biochemistry & Cell Biology and director of the Airola Lab. "Using this structure also helps us understand how the protein interacts with membranes, which is key to regulating its activity and the production of triglycerides."
The team used X-ray crystallography, mass spectrometry and biochemistry to visualize the structure, which represents the active state of a lipin enzyme during the production of triglycerides, as well as its other functions namely lipoprotein assembly and cellular signaling.
Valerie Khayyo, a Stony Brook graduate student in the Biochemistry & Structural Biology Program and first author of the study, added that the structure enables researchers to understand and see specific mutational changes in the amino acid building blocks of lipins that result in disease.
Khayyo noted that mutations in lipins are increasingly being identified in patients with muscle disorders as well, such as statin-induced myopathy and extreme cases in childhood rhabdomyolysis.
Overall, Airola says that the structure and study consolidates many previous observations into a unifying framework and sets the stage for scientists to resolve several remaining important questions concerning how lipins are regulated.
Journal information: Nature Communications
Provided by Stony Brook University