Scientists create triple-threat genetic toolkit for producing eco-friendly chemicals

Researchers have developed a triad of innovative tools to engineer low-pH-tolerant yeast Issatchenkia orientalis for production of valuable bioproducts from renewable biomass.
A paper published in Metabolic Engineering outlines the study's three-pronged approach and its importance to the field of sustainable chemical production.
The team was led by Mingfeng Cao, a Research Scientist in the University of Illinois at Urbana-Champaign's Department of Chemical and Biomolecular Engineering (ChBE). Cao works in ChBE Professor Huimin Zhao's lab at the Center for Advanced Bioenergy and Bioproducts Innovation (CABBI), a U.S. Department of Energy-funded Bioenergy Research Center (BRC). CABBI Postdoctoral Researcher Zia Fatma, also in Zhao's lab, is a coauthor of the study.
In an effort to decrease dependence on nonrenewable fossil fuels, scientists are exploring methods of producing valuable products from renewable biomass using organisms such as yeast. Generating economically competitive quantities of these products involves metabolic engineering—editing cells' genetic blueprint to produce greater quantities of desired substances.
Scientists previously engineered the conventional yeast Saccharomyces cerevisiae in this manner. A commonly used production host, S. cerevisiae's conduciveness to genetic manipulation makes it a prime candidate for metabolic engineering.
"In addition to conventional yeasts, we're discovering that nonconventional yeasts can prove to be just as high-performing," Cao said. "The more organisms we can engineer for valuable compound production, the more opportunities we will have in order to make chemical production economically and environmentally sustainable."
The study by Cao and Zhao focuses on the yeast strain I. orientalis, notable for its ability to thrive in low-pH environments. Scientists believe that this tolerance for extreme acidity positions I. orientalis as a robust organic acid producer.
Until now, the viability of I. orientalis for chemical production has remained less explored due to a shortage of genetic manipulation strategies. CABBI scientists remedied this knowledge gap by inventing three key tools.
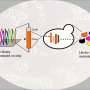
Previously, the research team had identified a genetic tool involving plasmids, DNA molecules necessary for genetic manipulation and reproduction. To edit an organism's genome, plasmids must be able to replicate stably. The team proved that an autonomously replicating DNA sequence (ARS) from Saccharomyces cerevisiae can be used to stabilize plasmid replication in I. orientalis, but only to an extent; on its own, the addition is insufficient.
Achieving stability and making I. orientalis receptive to CRISPR/CAS9 gene editing systems requires that the ARS is augmented with a functional centromere (CEN), the portion of a DNA sequence responsible for stabilizing cell division. The first novel genetic tool identified in this study is a functional CEN sequence in the I. orientalis genome. Because a CEN-ARS pairing is necessary for stable plasmid division, this discovery facilitates genetic engineering of I. orientalis for high-value, high-quantity compound production.
In addition to isolating a CEN sequence in I. orientalis, the research team identified constitutive promoters and terminators (components of a DNA strand that influence the way in which genes are expressed).
Finely tuned genetic expression—directly resulting from terminator and promoter behavior—is key to optimizing an organism for chemical production. By characterizing strong terminators and promoters in I. orientalis, researchers can engineer the particular expressions necessary to produce chemicals at high levels.
The third tool CABBI researchers developed is a fast and efficient in vivo (meaning "within a living organism") DNA assembly method to construct a biochemical pathway in I. orientalis.
For processes in I. orientalis to be high-yielding and economically viable, cellular pathways—chains of actions and reactions within a cell that cause a desired product or response—must often be engineered.
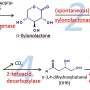
Taking a xylose utilization pathway originally designed for S. cerevisiae as an example, the research team used an in vivo DNA assembly method to introduce one such synthetic pathway in I. orientalis. As a result, I. orientalis could use xylose as a sole carbon source.
Though not fully optimized, this example demonstrated that a more sophisticated in vivo assembly method can be adopted for efficient construction of reliable biosynthetic pathways within I. orientalis, ultimately driving toward valuable compound production.
In conclusion, this study generated a robust set of three novel genetic tools to facilitate increased expression of target genes and aid in manufacturing of bioproducts such as organic acids from the yeast I. orientalis.
More information: Mingfeng Cao et al, A genetic toolbox for metabolic engineering of Issatchenkia orientalis, Metabolic Engineering (2020). DOI: 10.1016/j.ymben.2020.01.005
Provided by University of Illinois at Urbana-Champaign