February 20, 2020 report
High frequency of unwanted duplications in CRISPR-Cas9 edits

A team of researchers working at the University of Münster has found a high frequency of unwanted duplications during routine CRISPR-Cas9 genetic insertions in mice. In their paper published in the journal Science Advances, the group describes how they uncovered the unwanted duplications and give a warning to other researchers.
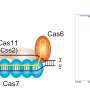
CRISPR-Cas9 is a gene editing technique developed over the past decade. It snips out undesired sections of a genome and inserts new sections. Much research has been conducted to test the procedure in the hopes that it can one day be used to repair genetic defects that lead to diseases. Progress toward that goal has been stymied by reports of off-target edits, which has led to new research aimed at preventing them. In this new effort, the researchers have found that the technique can also result in high numbers of unwanted duplications.
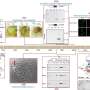
The finding was accidental, as they were studying encoding of a calcium binding protein by the gene S100A8 as a part of an immunology effort. To that end, they used CRISPR-Cas9 to disable the gene from expressing the protein—a form of knockout editing. They followed up their effort by testing the resulting genes to ensure things had gone as planned using a standard PCR test followed by a more specialized PCR test. The results showed that the editing had been successful in only two of the edits, which surprised them. The team then mated one of the mice with the edited gene with a wild mouse as part of an effort to understand why their success rate was so low. Testing of the offspring using a specialized type of PCR showed that seven of the mice had the edited genes—and the rest had unwanted duplicates.
Alarmed by their findings, the researchers carried out a second study in which they edited a different mouse gene. Specialized testing showed that out of 50 animals tested, 30 had multiple unwanted copies of a section of genome that had been inserted as part of CRISPR-Cas9 editing. And once again, the standard PCR tests failed to find them.
The researchers suggest their experience indicates that there are likely unwanted copies of gene insertions in prior work by others that has gone undocumented. They further suggest that researchers in the future use the more specialized testing techniques.
More information: Boris V. Skryabin et al. Pervasive head-to-tail insertions of DNA templates mask desired CRISPR-Cas9–mediated genome editing events, Science Advances (2020). DOI: 10.1126/sciadv.aax2941
Journal information: Science Advances
© 2020 Science X Network