August 5, 2016 report
Variation on CRISPR-Cas9 system offers editing without clipping both strands of DNA

(Phys.org)—A team of researchers with members from several institutions in Japan has developed a new way to edit genes that involves cutting just one strand of DNA rather than both of them, as is normal for CRISPR-Cas9. In their paper published in the journal Science, the team describes the technique and why they believe it is better for certain applications.
The CRISPR-Cas9 gene editing technique has made headlines over the past few years because it allows for relatively easy DNA substitutions. It involves introducing a bacterium into a cell along with a template—the bacterium cleaves the DNA strands at a certain position and then the cell makes repairs using the template—the result is a helix that has been edited to suit particular needs. But sometimes things go wrong because other cell mechanisms get involved, which can lead to unwanted deletions or insertions. To get around that problem, at least in part, the researchers have combined Cas9 with a deaminase (an enzyme that is capable of removing an amino group from a compound) from a sea lamprey that allows for single-stranded editing.
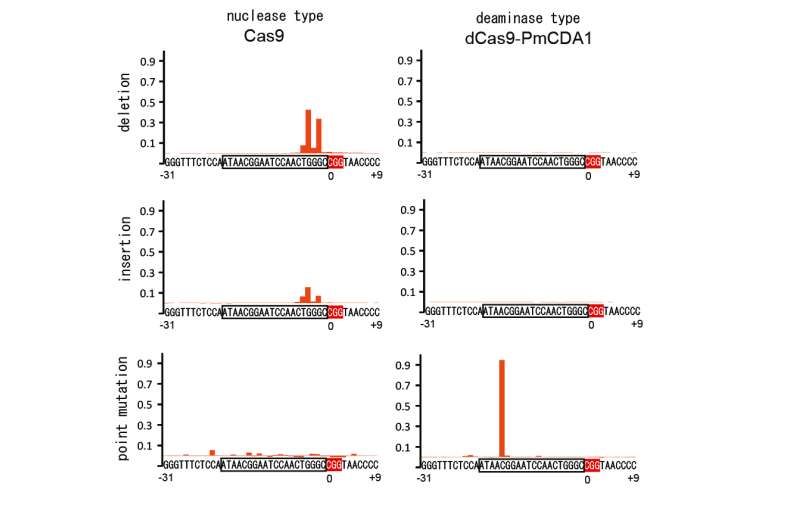
The researchers took two routes to the same objective, one using a nuclease-dead version of Cas9, which rendered it unable to cleave strands at all, and another using a nickase Cas9 that was able to create what they describe as a nick—clipping just one strand. In both approaches, the resulting Cas9 was fused with an activation-induced cytidine deaminase, which allowed for cutting just one strand and then doing its substitution work—all without the need for a template. After much testing, the team concluded that using the nickase Cas9 approach was more efficient. They also report that they were also able to make changes to two genes at the same time and that by adding another enzyme, they were able to increase the efficiency of the technique.
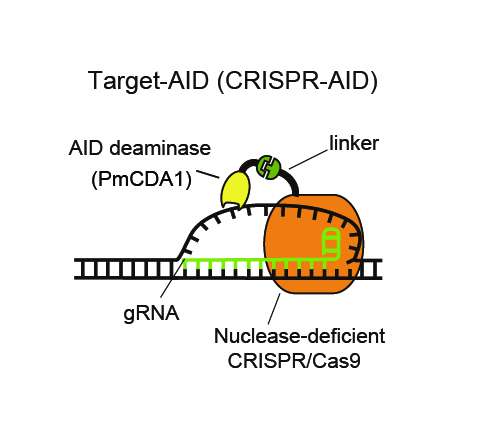
The team tested the technique on a yeast sample (because it did not have an internal gene altering system of its own) and found that it resulted in fewer collateral mutations than regular CRISPR-Cas9. They suggest the technique might prove useful in gene editing research with other bacteria and cell types that are particularly prone to mutations.
More information: K. Nishida et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems, Science (2016). DOI: 10.1126/science.aaf8729
Abstract
The generation of genetic variation (somatic hypermutation) is an essential process for the adaptive immune system in vertebrates. We demonstrate the targeted single-nucleotide substitution of DNA using hybrid vertebrate and bacterial immune systems components. Nuclease-deficient type II CRISPR/Cas9 and the activation-induced cytidine deaminase (AID) ortholog PmCDA1 were engineered to form a synthetic complex (Target-AID) that performs highly efficient target-specific mutagenesis. Specific point mutation was induced majorly at cytidines within the target range of five bases. The toxicity associated with the nuclease-based CRISPR/Cas9 system was greatly reduced. Although combination of nickase Cas9(D10A) and the deaminase was highly effective in yeasts, it also induced insertion and deletion (indel) in mammalian cells. Use of uracil DNA glycosylase inhibitor (UGI) suppressed the indel formation and improved the efficiency.
Journal information: Science
© 2016 Phys.org