April 21, 2016 report
New gene-editing technique improves on CRISPR allowing editing of single DNA letters
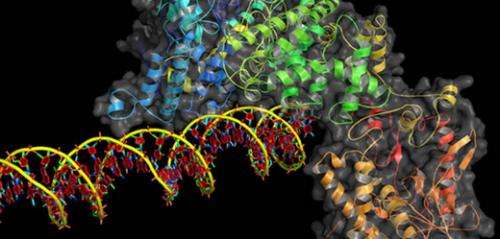
(Phys.org)—A team of researchers affiliated with Harvard University and the Howard Hughes Medical Institute has announced the development of a gene-editing system that improves on the performance of CRISPR/Cas9 by allowing for editing single DNA letters. In their paper published in the journal Nature, the team describes the technique, how it works and their hopes for its use in preventing disease sometime in the distant future.
As the researchers note, CRISPR/Cas9 was a big breakthrough in genetic engineering research, but it has suffered from one serious flaw—because it cuts both strands of DNA, errors creep in, making the results less than optimal. Also, the technique does not allow for making changes (point mutations) to a single DNA letter. In this new effort, the researchers describe a technique they have developed that improves on CRISPR/Cas9 in a way that does allow for fixing point mutations.
Normally, CRISPR/Cas9 works by using the Cas9 enzyme to target a specific spot on the genome—once that spot is identified, both strands are cut, which allows for insertion of DNA information; but cells react to the snipping as well, often putting in or taking out DNA letters which can alter the genome in undesirable ways. The new technique involves disabling the Cas9 enzyme so that when it finds the desired spot on the genome, it does not cut it, instead it carries with it another enzyme that is capable of changing one DNA letter to another. The new approach was found to work successfully 44 percent of the time on DNA that had been isolated, much better than the regular CRISPR/Cas9 technique. The group also tried adding a different protein to the bundle that was able to remove U basses and found it bumped the success rate up to 75 percent in mouse cells that were associated with Alzheimer's disease.
In the same journal issue, Nature has published two other articles related to gene editing, both of which involved teams looking into the possibility of replacing the Cas9 enzyme with another enzyme called Cpf1. One team in China described how Cpf1 changes shape when it is bound to guide RNA. The other team with members from Sweden and Germany described a technique for using Cpf1 to cut both RNA and DNA, opening up greater options for editing.
More information: Alexis C. Komor et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage, Nature (2016). DOI: 10.1038/nature17946
Abstract
Current genome-editing technologies introduce double-stranded (ds) DNA breaks at a target locus as the first step to gene correction. Although most genetic diseases arise from point mutations, current approaches to point mutation correction are inefficient and typically induce an abundance of random insertions and deletions (indels) at the target locus resulting from the cellular response to dsDNA breaks. Here we report the development of 'base editing', a new approach to genome editing that enables the direct, irreversible conversion of one target DNA base into another in a programmable manner, without requiring dsDNA backbone cleavage or a donor template. We engineered fusions of CRISPR/Cas9 and a cytidine deaminase enzyme that retain the ability to be programmed with a guide RNA, do not induce dsDNA breaks, and mediate the direct conversion of cytidine to uridine, thereby effecting a C→T (or G→A) substitution. The resulting 'base editors' convert cytidines within a window of approximately five nucleotides, and can efficiently correct a variety of point mutations relevant to human disease. In four transformed human and murine cell lines, second- and third-generation base editors that fuse uracil glycosylase inhibitor, and that use a Cas9 nickase targeting the non-edited strand, manipulate the cellular DNA repair response to favour desired base-editing outcomes, resulting in permanent correction of ~15–75% of total cellular DNA with minimal (typically ≤1%) indel formation. Base editing expands the scope and efficiency of genome editing of point mutations.
Ines Fonfara et al. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA, Nature (2016). DOI: 10.1038/nature17945
De Dong et al. The crystal structure of Cpf1 in complex with CRISPR RNA, Nature (2016). DOI: 10.1038/nature17944
Journal information: Nature
© 2016 Phys.org